Aim: To understand physiological and pathological roles of layilin in synovial fibroblasts, we comprehensively investigated protein profile changes caused by layilin-silencing. Methods: Immortalized human synovial membrane fibroblasts (HSFs) were transfected with siRNA for layilin (siL) in TNF-α-treated and -non-treated conditions. Proteins affected by siL were comprehensively detected by 2-dimensional fluorescence difference gel electrophoresis (2D-DIGE). Proteins of interest were identified by mass spectrometry. Results: In the 2D-DIGE analysis, intensity of approximately one-fifth of the detected protein spots was significantly changed by siL both in the TNF-α-treated and –non-treated conditions (239/1092 spots and 201/1092 spots, respectively). Proteins were identified in 24 out of 87 protein spots with ±1.3-fold or greater intensity changes by siL. 15 (62.5%) of the 24 protein spots were assigned to epithelial-mesenchymal transition (EMT)-related proteins. Conclusion: Our data suggest that layilin is deeply involved in the regulation of EMT-related proteins. Functions of layilin in synovial fibroblasts should be investigated in the context of EMT, although synovial fibroblasts are not epithelial cells. Furthermore, the EMT-related proteins affected by layilin may be involved in the pathogenesis of rheumatoid arthritis, which was characterized by proliferation and invasion of synovial fibroblasts.
Layilin, synovial membrane fibroblasts, TNF-α
Layilin is a type 1 transmembrane protein with a C-type lectin-like motif [1]. Layilin has been reported to be a receptor for hyaluronic acid [1] and also reported to bind to cytoskeletal proteins such as talin and merlin to regulate cell adhesion [2-4]. In fact, several studies showed that down-regulation of layilin inhibited infiltration and metastasis of cancer cells [5, 6]. Recently, it has been reported that layilin is highly expressed in an exhausted subset of CD8+ T cells in liver cancer [7]. We have reported that renal tubular epithelial cells in patients with glomerulonephritis more strongly express layilin than those in healthy people [8]. Taking these reports together, layilin is expected to play pathophysiological roles in various diseases. However, physiological and pathological roles of layilin have been poorly understood.
Accordingly, roles of layilin in rheumatoid arthritis (RA) remain also unclear. RA, a chronic polyarthritis, is histologically characterized by proliferation and invasion of synovial fibroblasts and infiltration of inflammatory cells into the expanded synovia in affected joints. In RA, inflammatory cytokines like tumor necrosis factor-α (TNF-α) are known to play pathological roles [9]. In fact, blockage of the inflammatory cytokines is used as effective therapies in RA [10]. Recently, Snail, a transcriptional factor, has been reported to regulate TNF-α-mediated activation of synovial fibroblasts in rheumatoid joints. We have reported that TNF-α up-regulates expression of layilin in human chondrocytes and also in a human clear cell renal carcinoma cell line of KMRC-1 [8, 11]. We have demonstrated that layilin is involved in TNF-α-mediated epithelial-mesenchymal transformation (EMT) of KMRC-1 [8]. We thus speculate that layilin is involved in the TNF-α-mediated inflammatory processes in RA. However, physiological roles of layilin as well as pathological roles in synovial fibroblasts have been poorly understood. To promote understanding of roles of layilin in synovial fibroblasts, we here comprehensively investigated effects of layilin-silencing on human synovial membrane fibroblasts (HSFs) in the TNF-α-treated and -non-treated conditions using proteomics.
Cell culture
Immortalized human synovial membrane fibroblasts (HSFs) were purchased from Applied Biological Materials Inc. (abm, Richmond, Canada). HSFs were cultured in a Prigrow III medium (abm) supplemented with 10% fetal bovine serum (FBS, Wako, Japan), 100 units/ml penicillin, and 100 μg/ml streptomycin (Sigma-Aldrich, St. Louis, MO, USA). The cells were cultured at 37°C in 5% CO2.
Treatment of HSFs with siRNA and TNF-α
HSFs were transfected with 200 pmol/f100 mm dish of siRNA for human LAYN (nucleotides 462-486 (GGGAGGGACACAGAGGCCTTGTTAT) in NM_001258390.1, Invitrogen, Carlsbad, CA, USA)) using Lipofectamine RNAiMAX (Invitrogen). As a negative control, the cells were transfected with the same doses of control siRNA (Stealth™ RNAi Negative Control Medium GC Duplex, Invitrogen). After 24 hours, the HSFs were further cultured in the starvation condition of a Prigrow III medium supplemented with 1% FBS, 100 units/ml penicillin, and 100 μg/ml streptomycin for 24 hours. Then the cells were incubated with or without 10 ng/ml human TNF-α (PROSPEC, Ness Ziona, Israel) for 24 hours. Proteins were extracted from the cells into a lysis buffer (7M urea, 2M thiourea, 4% CHAPS). The protein samples were stored at -80 °C until use.
RNA extraction, reverse transcription, and quantitative polymerase chain reaction (qPCR)
Total RNA, purified from HSFs using RNeasy® (Qiagen, Venlo, the Netherlands), was converted to cDNA by High Capacity cDNA Reverse Transcription Kits® (Life Technologies, Rockville, MD, USA), according to the manufacturer’s instructions.
qPCR was performed by ABI Prism 7000 Sequence Detection System (Applied Biosystems, Foster city, CA, USA). To measure mRNA for layilin and glyceraldehyde 3-phosphate dehydrogenase (GAPDH), a mixture of 2 µg cDNA, 300 nM each of forward and reverse primers, and Power SYBR® Master Mix (Applied Biosystems) was subjected to qPCR. Nucleotide sequences of the primers are as follows: layilin: 5’ -CACAGCCTGCCAGGACCTTTA and 5’ -TGCACCGGTCATCATTCCA, and GAPDH: 5’-TGGTATGGTGGAAGGACTCA and 5’-ATGCCAGTGAGCTTCCCGTT. The thermal cycle condition was as follows: 95°C for 10 min, followed by 40 cycles of 95°C for 15 sec and 60°C for 60 sec.
2-dimensional fluorescence difference gel electrophoresis (2D-DIGE)
The extracted proteins were separated by 2D-DIGE as described previously [12, 13]. Briefly, each of the protein samples was labeled with Cyanine dye 5 (Cy5, Cy Dye DIGE Saturation dye; GE Healthcare, Buckinghamshire, UK). An internal standard sample was prepared by mixing an equal amount of each of the samples. The standard sample was then labeled with Cyanine dye3 (Cy3, GE Healthcare). Then, each of the Cy5-labeled protein samples (2.5 µg) were mixed with the Cy3-labeled standard sample (2.5 µg). The mixture was applied onto a non-liner isoelectric focusing (IEF) gel (24 cm in width, pH 3-11, GE Healthcare). Subsequently, proteins separated by IEF were further separated by 12.5 % sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE). Then, the separated protein spots were scanned using an image analyzer (Typhoon 9400 Imager, GE Healthcare). On each protein spot, Cy5- and Cy3-fluorescence intensity was separately quantified by the Progenesis program (PerkinElmer, MA, US) and the Cy5-fluorescence intensity was normalized by Cy3-fluorescence intensity. The normalized Cy5-intensity was used for the quantitative comparison among the samples.
Protein identification
Protein spots of interest, excised from the 2D gels, were digested by trypsin as described previously [14]. Produced peptides were subjected to matrix-assisted laser desorption ionization-time of flight/time of flight mass spectrometry (MALDI-TOF/TOF-MS) (Ultraflex, BrukerDaltonics, Bremen, Germany). Obtained peptide masses were compiled to allow searches of the National Center for Biotechnology Information (NCBI) protein database using the Mascot software program (Matrix Science, London, UK).
Statistical analysis
Statistical significance was calculated using Student’s t-test.
Detection of proteins affected by layilin-silencing
We here comprehensively investigated changes of protein profiles of HSFs caused by siRNA for layilin in TNF-α-treated and non-treated conditions, using 2D-DIGE and mass spectrometry. First, HSFs were transfected with siRNA for human layilin (siL) or control siRNA (siC). In part, the cells were further treated with TNF-α for 24 hours. Then the cultured cell samples were subjected to 2D-DIGE. Prior to the experiments, the efficacy of layilin-silencing by siL was checked by qPCR. As a result, expression of layilin was suppressed down to approximately 7% (Figure 1), indicating that layilin-silencing was enough for the analysis.
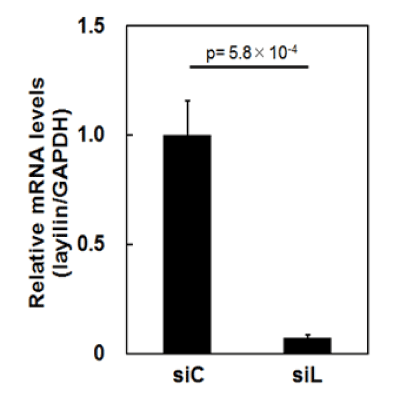
Figure 1. Silencing of layilin by siRNA in HSFs
HSFs were transfected with layilin or control siRNA (siL or siC, respectively). Then, RNA extracted from the cells was used for quantitative real time-PCR to estimate amounts of mRNA for layilin and GAPDH as a control. The measured mRNA levels of layilin were corrected using those of GAPDH. The average of the corrected mRNA levels in the cells transfected with siC was defined as 1.0. Mean values with SD are shown.
Comprehensive analysis of proteins affected by layilin-silencing in the TNF-α- treated or non-treated condition by 2D-DIGE
By the 2D-DIGE analysis, in total, 1092 protein spots were detected on the 2D-gels (Figure 2A). In the condition without TNF-α, 239 (21.9%) out of the 1092 spots showed significantly different intensity between the siL and siC conditions (Table 1). Among the 239 spots, 29 spots showed 1.3-fold or higher intensity in the siL condition than in the siC condition and 24 spots showed 1/1.3-fold or lower intensity in the siL condition than in the siC condition (Table 1). Focusing on the 29 and 24 protein spots, we then compared intensity between the siL and siC conditions under the TNF-α-treated condition (Table 1). As a result, 17 out of the 29 spots showed significantly higher intensity in the siL condition than in the siC condition even under the TNF-α-treated condition (Category 1). The other 12 spots did not show significant difference between the siL and siC conditions under the TNF-α-treated condition (Category 2). No spots showed significantly lower intensity in the siL condition than in the siC condition (Category 3). Similarly, 2 out of the 24 spots showed significantly lower intensity in the siL condition than in the siC condition even under the TNF-α-treated condition (Category 6). The other 22 spots did not show significant difference between the siL and siC conditions (Category 5). No spots showed significantly higher intensity in the siL condition than in the siC condition (Category 4). Proteins of the spots in Categories 1 and 6 are thought to be affected by layilin, regardless of presence or absence of TNF-α. Proteins of the spots in Categories 2 and 5 are thought to be affected by layilin only in the TNF-α-non-treated condition.
Table 1. Numbers of protein spots intensity of which was significantly altered by layilin-silencing in the TNF-α-non-treated condition
Total number of protein spots |
TNF-a (-) |
TNF-a (+) |
Category in this study |
Number of spots with different intensity (p<0.05) |
Number of spots classified by siL/siC |
1092 |
239 |
siL/siC≥1.3 |
29 |
↑ |
17 |
1 |
→ |
12 |
2 |
↓ |
0 |
3 |
siL/siC≤-1.3 |
24 |
↑ |
0 |
4 |
→ |
22 |
5 |
↓ |
2 |
6 |
On the other hand, in the TNF-α-treated condition, 201 (18.4%) out of the 1092 spots showed significantly different intensity between the siL and siC conditions (Table 2). Among the 201 spots, 14 spots showed 1.3-fold or higher intensity in the siL condition than in the siC condition and 20 spots showed 1/1.3-fold or lower intensity in the siL condition than in the siC condition (Table 2). Focusing on the 14 and 20 protein spots, we then compared intensity between the siL and siC conditions under the TNF-α-non-treated condition (Table 2). As a result, 5 out of the 14 spots showed significantly higher intensity in the siL condition than in the siC condition even under the TNF-α-non-treated condition (Category 7). The other 9 spots did not show significant difference between the siL and siC conditions under the TNF-α-non-treated condition (Category 8). No spots showed significantly lower intensity in the siL condition than in the siC condition (Category 9). Similarly, 4 out of the 20 spots showed significantly lower intensity in the siL condition than in the siC condition even under the TNF-α-non-treated condition (Category 12). The other 16 spots did not show significant difference between the siL and siC conditions under the TNF-α-non-treated condition (Category 11). No spots showed significantly higher intensity in the siL condition than in the siC condition (Category 10). Proteins of the spots in Categories 7 and 12 are thought to be affected by layilin, regardless of presence or absence of TNF-α. Proteins of the spots in Categories 8 and 11 are thought to be affected by layilin only in the TNF-α-non-treated condition.
Table 2. Numbers of protein spots intensity of which was significantly altered by layilin-silencing in the TNF-α-treated condition
Total number of protein spots |
TNF-a (+) |
TNF-a (-) |
Category in this study |
Number of spots with different intensity (p<0.05) |
Number of spots classified by siL/siC (p<0.05) |
1092 |
201 |
siL/siC≥1.3 |
14 |
↑ |
5 |
7 |
→ |
9 |
8 |
↓ |
0 |
9 |
siL/siC≤-1.3 |
20 |
↑ |
0 |
10 |
→ |
16 |
11 |
↓ |
4 |
12 |
Identification of proteins that are affected by layilin-silencing
Next, we tried to identify proteins of the 53 protein spots affected by layilin-silencing in the TNF-α-non-treated condition (29 up-regulated and 24 down-regulated, Table1), using MALDI-TOF-TOF mass spectrometry. As a result, 15 out of the 53 protein spots were identified (Figure 2B, Table 3).
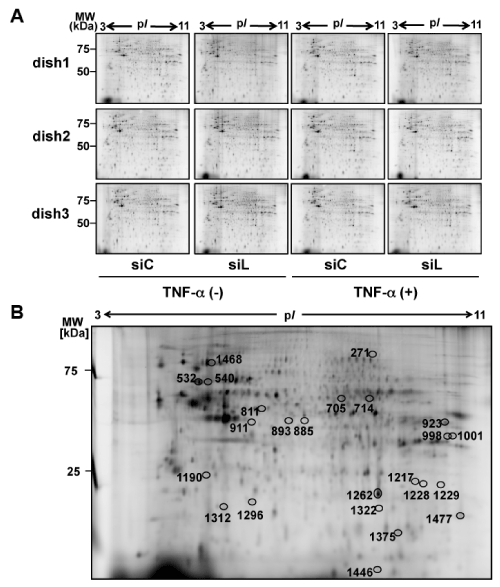
Figure 2. 2D-DIGE analysis of effects of layilin-silencing on HSFs
(A) HSFs were transfected with siRNA for layilin (siL) or control siRNA (siC). Then the cells were cultured for 24 hours and then subjected to 24 hours starvation. Next, a part of the cultured cells were further treated with 10 ng/ml TNF-α for 24 hours. Proteins, extracted from the HSFs were subjected to 2D-DIGE (n=3). MW, molecular weight.
(B) Protein spots intensity of which was significantly altered by siL were subjected to protein identification by MALDI-TOF-TOF/MS. Locations of the identified spots on the 2D gel are shown by circles with spot numbers.
Proteins of the spots in Category 1, which were thought to be down-regulated by layilin regardless of presence or absence of TNF-α, were identified as follows: ID1217: superoxide dismutase [Mn] mitochondrial (SODM), ID893: calcium-binding mitochondrial carrier protein SCaMC-1 (SCMC1), ID923: serpin H1 (SERPH), ID532: 78kDa glucose-regulated protein (GRP78), ID1001: heterogeneous nuclear ribonucleoproteins A2/B1 (ROA2), and ID1477: peptidyl-prolyl cis-trans isomerase B. Proteins of the spots in Category 2, which were thought to be down-regulated by layilin only under the TNF-α-non-treated condition, were identified as follows: ID1262: SODM, ID998: ROA2, ID714: nicotinamide phosphoribosyltransferase (NAMPT), ID911: keratin, type II cytoskeletal 1 (K2C1), and ID885: SCMC1. Proteins of the spots in Category 5, which were thought to be up-regulated by layilin only under the TNF-α-non-treated condition, were identified as follows: ID540: GRP78, ID271: growth /differentiation factor 10 (GDF10), ID811: glutathione synthetase (GSHB), and ID1190: melanoma-associated antigen B3 (MAGB3).
Similarly, we tried to identify proteins of the 34 protein spots affected by layilin-silencing under the TNF-α-treated condition (14 up-regulated and 20 down-regulated, Table 4). As a result, 10 out of the 34 proteins spots were identified (Figure 2B, Table 4).
Proteins of the spots in Category 7, which were thought to be down-regulated by layilin regardless of presence or absence of TNF-α, were identified as follows: ID1217: SODM and ID1312: peroxyiredoxin-2. Proteins of the spots in Category 8, which were thought to be down-regulated by layilin only under the TNF-α-treated condition, were identified as follows: IDs1228 and 1229: protein NipSnap homolog 3A (NPS3A). Proteins of the spots in Category 11, which were thought to be up-regulated by layilin only under the TNF-α-treated condition, were identified as follows: ID1468: vimentin, ID705: cdc42-interacting protein 4 (CIP4), ID1322: peroxisomal bifunctional enzyme (ECHP), ID1375: zinc finger protein 350 (ZN350), ID1446: cystatin-B (CYTB ), and ID1296: peroxyiredoxin-2 (PRDX2).
We here comprehensively investigated effects of layilin-silencing on HSFs under TNF-α-treated and -non-treated conditions using proteomics. We found that layilin-silencing significantly changed the expression of approximately one-fifth of the detected protein spots both in the two conditions. The 53 spots that showed ±1.3-fold or greater intensity changes in the TNF-α-non-treated condition were further classified into Categories 1-6, according to the changes by layilin-silencing under the TNF-α-treated condition. Similarly, the 34 protein spots that showed ±1.3-fold or greater intensity changes in the TNF-α-treated condition were further classified into Categories 7-12, according to the changes by layilin-silencing under the TNF-α-non-treated condition.
2021 Copyright OAT. All rights reserv
The protein spots of Categories 1 and 7 and Categories 6 and 12 are thought to contain proteins that regulated by layilin independently from TNF-α signaling. Here, 7 proteins were identified in Categories 1 and 7. Interestingly, 5 out of the 7 proteins were reported to be related with EMT (ID1217: SODM、ID923: SERPH、ID532: GRP78、ID1001: ROA2、ID1312: Peroxiredoxin-2.). Specifically, SODM, an enzyme that catalyzes the superoxide radical, has been reported to be up-regulated during EMT induced by TNF-α and to enhance expression of CD44, which are strongly expressed on highly malignant cancer cells [15]. SERPH, one of heat shock proteins, has been reported to be increased during cyclosporine-induced EMT of renal epithelial cells in mice [16]. GRP78, one of heat shock proteins, has been reported to increase fibronectin and integrin β1 to enhance adhesiveness of colon cancer cells [17]. Moreover, expression of GRP78 has been reported to be increased at EMT of human colon adenocarcinoma [17]. ROA2, a nuclear ribonucleoprotein, has been reported to be highly expressed in lung cancer cells [18]. Silencing of ROA2 have been reported to up-regulate E-cadherin and down-regulate transcriptional repressors of E-cadherin [18], thereby, ROA2 is thought to promote EMT. Collectively, these 4 proteins are thought to be involved in induction and/or promotion of EMT. The up-regulation of these 4 proteins by layilin-silencing observed here indicates that layilin constitutively suppresses these 4 proteins. On the other hand, PRDX2 has been reported to inhibit TNF-α-induced EMT in colon cancer cells [19]. The up-regulation of PRDX2 by layilin-silencing observed here indicates that layilin constitutively suppresses PRDX2.
The protein spots of Categories 2 and 5 are thought to contain proteins that appear to be affected by layilin only under the TNF-α-non-treated condition. These proteins may be regulated oppositely by TNF-α and layilin-silencing. Alternatively, if TNF-α down-regulates layilin, layilin-silencing would not actually affect the proteins under the TNF-α-treated condition. In Categories 2 and 5, 9 proteins were identified. Five out of the 9 proteins, i.e., ID1262: SODM、ID998: ROA2、ID714: NAMPT、ID271: GDF10、ID811: GSHB, were reported to be related to EMT. Specifically, SODM identified from ID1262 and ROA2 identified from ID998 were also identified from the spots of ID1217 and ID1001 in Category 1, respectively. This indicates existence of different post-translational modifications on SODM and ROA2. As above described, both SODM and ROA2 have been reported to induce or promote EMT. NAMPT, a rate-limiting enzyme of NAD+ synthesis, has been recently reported to promote EMT, independently from its enzyme activity [20]. GDF10, a member of the TGF-β super family, has been reported to suppress EMT in oral squamous cell carcinoma [21]. On GSHB of a glutathione synthase, decrease of GSHB has been reported to promote EMT, resulting in fibrosis [22]. Thus, all the 5 EMT-related proteins, SODM, ROA2, NAMPT, GDF10, and GSHB are thought to induce or promote EMT.
The protein spots of Categories 8 and 11 are thought to contain proteins that are regulated by layilin in the TNF-α signaling pathway. Out of the 8 identified proteins in Categories 8 and 11, 4 proteins (50%), CIP4, CYTB, PRDX2, and VIME, were reported to be related to EMT. Since all the 4 proteins were down-regulated by layilin-silencing, these 4 proteins were thought to be up-regulated by TNF-α via layilin. CYTB and PRDX2 have been reported to be suppressed during EMT or to suppress EMT [19, 23]. On the other hand, CIP4 and VIME have been reported to be increased during EMT or to promote EMT [24, 25].
Collectively, 15 (62.5%) out of the 24 identified proteins spots were found to be EMT-related ones. Among the 15 proteins, 10 proteins are related to induction or promotion of EMT as above discussed. Here, layilin-silencing up-regulated 7 out of the 10 proteins, and at the same time, down-regulated the remaining 3 proteins. Similarly, the remaining 5 out of the 16 proteins are related to suppression of EMT as above discussed. Here, layilin-silencing down-regulated 4 out of the 5 proteins, and at the same time, up-regulated the remaining 1 protein. Thereby, overall effects of layilin on EMT cannot be easily determined at present, although layilin is thought to be involved in EMT. Further studies should be needed to elucidate precise roles of layilin in EMT and EMT-like changes.
Again, 15 (62.5%) out of the 24 identified protein were found to be EMT-related ones. This result suggests EMT-related roles of layilin in the pathogenesis of RA. Synovial fibroblasts in patients with RA are known to possess high growth potential and high invasiveness into cartilage and bone [26]. Recently, it has been reported that these character changes of synovial fibroblasts in RA are associated with up-regulation of various EMT markers [27, 28]. Thereby, mechanisms that cause EMT are probably involved in the character changes of synovial fibroblasts, although synovial fibroblasts are not epithelial cells.
In conclusion, this study suggests that layilin is deeply involved in the regulation of EMT-related proteins in synovial fibroblasts. Functions of layilin in synovial fibroblasts in RA as well as in healthy conditions should be investigated from the viewpoint of EMT.
We thank Ms. Yokoyama MK and Ms. Sawada Y for their technical assistance.
None.
- Bono P, Rubin K, Higgins JM, Hynes RO (2001) Layilin, a novel integral membrane protein, is a hyaluronan receptor. Mol Biol Cell 12(4): 891-900. [Crossref]
- Borowsky ML, Hynes RO (1998) Layilin, a novel talin-binding transmembrane protein homologous with C-type lectins, is localized in membrane ruffles. J Cell Biol 143(2): 429-442. [Crossref]
- Wegener KL, Basran J, Bagshaw CR, Campbell ID, Roberts GC, et al. (2008) Structural basis for the interaction between the cytoplasmic domain of the hyaluronate receptor layilin and the talin F3 subdomain. J Mol Biol 382(1): 112-126. [Crossref]
- Scoles DR. (2008) The merlin interacting proteins reveal multiple targets for NF2 therapy. Biochim Biophys Acta 1785(1): 32-54. [Crossref]
- Zhuo W, Wang Y, Chen Z (2006) Inhibitory effects of shRNA targeting layilin on adhesion and invasion behavior of human lung adenocarcinoma A549 cells induced by hyaluronan in vitro. Zhongguo Fei Ai Za Zhi 9(3): 236-240. [Crossref]
- Chen Z, Zhuo W, Wang Y, Ao X, An J (2008) Down-regulation of layilin, a novel hyaluronan receptor, via RNA interference, inhibits invasion and lymphatic metastasis of human lung A549 cells. Biotechnol Appl Biochem 50(Pt 2): 89-96. [Crossref]
- Zheng C, Zheng L, Yoo JK, Guo H, Zhang Y, et al. (2017) Landscape of Infiltrating T Cells in Liver Cancer Revealed by Single-Cell Sequencing. Cell 169: 1342-1356. [Crossref]
- Adachi T, Arito M, Suematsu N, Kamijo-Ikemori A, Omoteyama K, et al. (2015) Roles of layilin in TNF-a-induced epithelial-mesenchymal transformation of renal tubular epithelial cells. Biochem Biophys Res Commun 467(1): 63-69. [Crossref]
- Brennan FM, Maini RN, Feldmann M (1992) TNF alpha--a pivotal role in rheumatoid arthritis? Br J Rheumatol 31: 293-298. [Crossref]
- Maini RN, Feldmann M (2002) How does infliximab work in rheumatoid arthritis? Arthritis Res 4 Suppl 2: S22-28. [Crossref]
- Asano K, Arito M, Kurokawa MS, Omoteyama K, Okamoto K, et al. (2014) Secretion of inflammatory factors from chondrocytes by layilin signaling. Biochem Biophys Res Commun 452: 85-90. [Crossref]
- Toda T, Ishijima Y, Matsushita H, Yoshida M, Kimura N (1994) Detection of thymopoietin-responsive proteins in nude mouse spleen cells by two-dimensional polyacrylamide gel electrophoresis and image processing. Electrophoresis 15: 984-987. [Crossref]
- Onodera H, Arito M, Sato T, Ito H, Hashimoto T, et al. (2013) Novel effects of edaravone on human brain microvascular endothelial cells revealed by a proteomic approach. Brain Res 1534: 87-94. [Crossref]
- Fujisawa H, Ohtani-Kaneko R, Naiki M, Okada T, Masuko K, et al. (2008) Involvement of post-translational modification of neuronal plasticity-related proteins in hyperalgesia revealed by a proteomic analysis. Proteomics 8: 1706-1719. [Crossref]
- Kinugasa H, Whelan KA, Tanaka K, Natsuizaka M, Long A, et al. (2015) Mitochondrial SOD2 regulates epithelial-mesenchymal transition and cell populations defined by differential CD44 expression. Oncogene 34(41): 5229-5239. [Crossref]
- Galichon P, Vittoz N, Xu-Dubois YC, Cornaire E, Vandermeersch S, et al. (2011) Epithelial phenotypic changes detect cyclosporine in vivo nephrotoxicity at a reversible stage. Transplantation 92(9): 993-998. [Crossref]
- Zhang L, Li Z, Fan Y, Li H, Li Z, et al. (2015) Overexpressed GRP78 affects EMT and cell-matrix adhesion via autocrine TGF-β/Smad2/3 signaling. Int J Biochem Cell Biol 64: 202-211. [Crossref]
- Tauler J, Zudaire E, Liu H, Shih J, Mulshine JL (2010) hnRNP A2/B1 modulates epithelial-mesenchymal transition in lung cancer cell lines. Cancer Res 70: 7137-7147. [Crossref]
- Feng J, Fu Z, Guo J, Lu W, Wen K, et al. (2014) Overexpression of peroxiredoxin 2 inhibits TGF-β1-induced epithelial-mesenchymal transition and cell migration in colorectal cancer. Mol Med Rep 10: 867-873. [Crossref]
- Soncini D, Caffa I, Zoppoli G, Cea M, Cagnetta A, et al. (2014) Nicotinamide phosphoribosyltransferase promotes epithelial-to-mesenchymal transition as a soluble factor independent of its enzymatic activity. J Biol Chem 289(49): 34189-34204. [Crossref]
- Cheng CW, Hsiao JR, Fan CC, Lo YK, Tzen CY, et al. (2016) Loss of GDF10/BMP3b as a prognostic marker collaborates with TGFBR3 to enhance chemotherapy resistance and epithelial-mesenchymal transition in oral squamous cell carcinoma. Mol Carcinog 55(5): 499-513. [Crossref]
- Liu RM, Gaston Pravia KA (2010) Oxidative stress and glutathione in TGF-beta-mediated fibrogenesis. Free Radic Biol Med 48: 1-15. [Crossref]
- Xiong Y, Ji W, Fei Y, Zhao Y, Wang L, et al. (2017) Cathepsin L is involved in X-ray-induced invasion and migration of human glioma U251 cells. Cell Signal 2017 29:181-191. [Crossref]
- Bai S, Zeng R, Zhou Q, Liao W, Zhang Y, et al. (2012) Cdc42-interacting protein-4 promotes TGF-β1-induced epithelial-mesenchymal transition and extracellular matrix deposition in renal proximal tubular epithelial cells. Int J Biol Sci 8(6):859-69. [Crossref]
- Kokkinos MI, Wafai R, Wong MK, Newgreen DF, Thompson EW, et al. (2007) Vimentin and epithelial-mesenchymal transition in human breast cancer--observations in vitro and in vivo. Cells Tissues Organs 185(1-3): 191-203. [Crossref]
- Firestein GS (1996) Invasive fibroblast-like synoviocytes in rheumatoid arthritis: passive responders or transformed aggressors? Arthritis Rheum 39: 1781–1790. [Crossref]
- Steenvoorden MM, Tolboom TC, van der Pluijm G, Löwik C, Visser CP, et al. (2006) Transition of healthy to diseased synovial tissue in rheumatoid arthritis is associated with gain of mesenchymal/fibrotic characteristics. Arthritis Res Ther 8(6):R165. [Crossref]
- Zvaifler NJ (2006) Relevance of the stroma and epithelial-mesenchymal transition (EMT) for the rheumatic diseases. Arthritis Res Ther 8: 210. [Crossref]


