Human genetic factors mechanisms are known to protect against malaria. However, the mechanism underlying the influence of human genetic variation on Plasmodium falciparum (P.falciparum) genetic diversity is still a research purpose. The aim of this study is to assess the effect of haemoglobin variants and Glucose-6-Phosphate-Dehydrogenase (G6PD) variation on the P. falciparum merozoite surface protein (msp1, msp2), and the glutamate rich protein (glurp) genetic diversity in children living in Burkina Faso (BF). A cross-sectional study was carried out at Banfora and Saponé health districts and 386 children less than five years were enrolled. DNA collected from each sample was extracted using chelex-100 method and then analyzed by a nested PCR of msp1, msp2 and glurp genes while the human genetic resistance factors background was assessed by RFLP-PCR. A total of 112 (29.02%) subjects were classified as abnormal haemoglobin and 38 (9.59%.) were identified as G6PD deficiency The distribution of the msp1 and msp2 allelic families was not different according to haemoglobin type (p=0.70 and 0.90 respectively) and G6PD type (p=0.89 and 0.82 respectively). The prevalence of the glurp gene was 93.00% and there was no statistical difference in its distribution according to the human factors (p=0.24 and p=0.95 for haemoglobin and G6PD types respectively). The analysis of the mean multiplicity of P. falciparum infection (MOI) based on haemoglobin variants showed msp1 with high values 2.96 and 3.12 for Normal haemoglobin and Abnormal haemoglobin respectively. However, according to the G6PD type, there were no differences of MOIs between normal G6PD and deficient G6PD carriers. The study showed the P. falciparum genetic diversity was not affected by human genetic factors based on the analysis of msp1, msp2 and glurp.
g6pd, glurp, haemoglobin, malaria, msp1, msp2
An estimated of 219 million cases of malaria occurred worldwide and most malaria cases in 2017 (92%) were living in the WHO African Region [1]. Despite efforts to combat malaria, the resistance of the parasite to antimalarial drugs has increased the macabre and economic burden of the plague in many endemic countries [2]. Significant decrease in malaria incidence rates globally and in the African region have been recorded and are the result of intensified monitoring and control measures and strategies [1]. However, these improvements remain fragile, owing to the relentless development of parasite resistance to antimalarial [3]. Of these, the development of an effective vaccine is the ultimate challenge for malaria control. The challenge is indeed more complex as Plasmodium is an extremely polymorphic species [4]. The diversity and antigenic variation of the parasite are two main factors responsible for the slow acquisition of malaria immunity [5]. The risk for an individual to manifest a certain type of symptoms depends on several factors such as genetic, immune, nutritional status; but also depends on the parasite himself [6]. In parallel, the existence of innate genetic factors and immunological mechanisms allows human to resist against malaria has been also described [7]. Indeed, several human genetic factors (abnormal haemoglobin, alpha and beta thalassemia, G6PD deficiency, TNFα mutation and HLA factor) were identified as conferring a certain degree of resistance to malaria [8,9] but there is not enough information on the influence of human genetic factors on the parasitic diversity of P. falciparum.
Given this, the aim of this study is to analyze the possible effect of human genetic resistance factors such as haemoglobin variant and G6PD deficiency on the genetic diversity of P. falciparum Merozoite Surface Proteins 1 and 2 (msp1 and msp2) and Glutamate-Rich Protein (glurp) genes in subjects with P. falciparum uncomplicated malaria, living in malaria endemic area of BF.
Study sites
The study was conducted in two areas covering the health district of Banfora and Saponé. Banfora health district is located in the Comoé province, at the southwestern part of Burkina Faso, at about 450 km from Ouagadougou, the capital city. Malaria is endemic and the transmission in that area is permanent with seasonal peaks during the rainy season from May to November. The health district of Saponé is located 50 Km south-west of Ouagadougou. Malaria transmission even is seasonal is short compared to the one of Banfora health district (June to October). According to the BF Ministry of Health, malaria incidence in the country was 364 ‰ and 413‰ in 2010 and 2013 respectively [10].
Study Population, design and period
A total of 386 children aged between 6 to 59 months were recruited. It was cross-sectional surveys with descriptive focus conducted from December 2010 to January 2013 in Saponé health district and from May to October 2011 in Banfora health district. The study was part of a clinical trial study, assessing the efficacy of two Artemisinin combination therapies (ACT). The inclusion criteria were as followed : 1) fever (axillary temperature ≥ 37.5°C) and/or a history of fever within the past 48 hours; 2) asexual P. falciparum mono-infection identified microscopically on blood smears with parasite density between 2000 and 200000 parasites/µl of blood; 3) no history of anti-malarial drug administration in the last two weeks; 4) no history of serious adverse effects to the study drugs (mefloquine, quinine, artesunate, chloroquine and sulphadoxine-pyrimethamine); 6) no evidence of a concomitant febrile illness; 7) no sign/symptoms of severe malaria as defined by WHO.
Samples collection
For each subject included in this study, physical examination and capillary blood samples on slides and filter papers were collected. Slides were used for the diagnosis of malaria parasites and the filter papers for the analysis of human genetic factors and the genetic diversity of P. falciparum.
Malaria Parasites Diagnosis by microscopy
P. falciparum parasitemia was microscopically diagnosed. Blood slides with thin and thick blood smears were prepared and stained with Giemsa (10%). The thick film was fixed with methanol after drying. The parasites were counted against 200 leukocytes and then extrapolated to parasites per microliter of blood. At least one hundred power film fields were examined before assigning a negative malaria diagnosis. The number of parasites per microliter of blood was calculated using the last full blood count of the patient or the theoretical value of 8000 leucocytes/μl. The Parasite Density (PD) was estimated using the following formula:
PD = N x 8000/X
With N = number of parasites counted and X = number of counted leucocytes or the value of the full blood count.
Two expert’s microscopists who read each blood slide were blinded from each other’s reading. All discordant readings were re-read by a third microscopist who was blinded from the previous results.
Blood spots samples and DNA extraction
Blood from finger prick spotted onto Whatman filter Papers (Whatman 3 mm, GE Healthcare, Pittsburg, USA), was labeled with patients’ study numbers, air-dried, and individually placed into plastic bag marked and containing a desiccant to protect against humidity. The bags were stored at room temperature until DNA extraction. Parasite DNA was extracted using Chelex methods [11]. Briefly, three pieces of filter paper was soaked overnight in a solution of 10% saponin in PBS and was subsequently washed in PBS. Thus, 50 μl of 20% Chelex® 100 solution (Bio-Rad Laboratories) were added to 1.5 ml micro centrifuge tube containing 3 fragments of filter paper sample. Then, 100 μl of sterile water were added and the micro centrifuge tube placed onto a heating block at 95-100°C for 10 minutes of incubation. During the incubation phase, the tube was gently whirled and returned to the heat block every two minutes. The samples were centrifuged twice and the final supernatant about 150 μl was conserved in a new labeled tube and stored at -20°C until it was used for the amplification reaction.
Haemoglobin genotyping
The haemoglobin in the β-chain of the globin gene at codon six was determined by using polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP). The PCR conditions were as follows: one (1) cycle of 5 min at 96°C, 30 cycles of 96°C for 30 secs, 60°C for 1 min, 72°C for 30 secs and 72°C for 7 min. DNA samples were amplified. The expected fragment length was 358 bases pairs (bp). The fragment obtained was digested for three hours at 37°C with MnlI for discrimination between HbAA (173 bp, 109 bp, and 60 bp), HbSS/HbCC and HbSC (173 bp, 109 bp, and 76 bp), HbAS/HbAC (173 bp, 109 bp, 76 bp and 60 bp). A second digestion with DdeI allowed for further discrimination for ambiguous results between HbSS (331 bp), HbCC (201 bp and 130 bp), HbSC (130 bp, 201 bp and 331 bp), HbAS (130 bp, 201 bp and 331 bp) and HbAC (201 bp and 130 bp). PCR products were analyzed by electrophoresis in a 1.5 % agarose gel.
G6PD genotyping
DNA was amplified and analyzed for the presence or absence of one of the common G6PD mutations G→A 202 using Polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP) method. PCR amplification was done using primers forward and reverse (Table 1). Details of the PCR-RFLP process are described elsewhere [12]. (Table 1).
Table 1. Primers sequences for haemoglobin type and G6PD amplification
Genes |
Primer name |
Allelic-specific primers |
Haemoglobin type |
Forward |
AGGAGCAGGGAGGGCAGGA |
Reverse |
TCCAAGGGTAGACCACCA GC |
G6PD type |
Forward |
GTGGCTGTTCCGGGATGGCCTTCG |
Reverse |
CTTGAAGAAGGGCTCACTCTGTTTG |
PCR amplification of the P. falciparum msp1, msp2 and glurp genes
Msp1 (region 2), msp2 (central region) and glurp were amplified by a nested PCR. Products obtained after the first PCR, were amplified using specific primers for msp1 where three distinct allelic families (K1, Mad20, and Ro33) have been described; two distinct families for msp2 (FC27 and 3D7) and glurp in accordance to the recommended genotyping protocol [13]. The sequence of the primers and the protocol of PCR were previously described in detail by Snounou et al [14]. The following oligonucleotide primers (Table 2) sequences were used. (Table 2).
Table 2.Primers sequences for msp1, msp2 and glurp.
Genes |
PCR round |
Primer name |
Allelic-specific primers |
msp1 |
Primary |
msp1-A
msp1-B |
AAGCTTTAGAAGATGCAGTATTGAC
ATTCATTAATTTCTTCATATCCATC |
Secondary |
K1-A
K1-B
Mad20-A
Mad20-B
Ro33-A
Ro33-B |
AAGAAATTACTACAAAAGGTGCAAGTG
AGAACAAGTCGAACAGCTGTTA
TGAATTATCTGAAGGATTTGTACGTCTTGA
GAGCAAGTCGAACAGCTGTTA
AGGATTTGCAGCACCTGGAGATCT
GAGCAAATACTCAAGTTGTTGCAAAGC |
msp2 |
Primary |
msp2-1
msp2-4 |
ATGAAGGTAATTAAAACATTGTCTATTATA
ATATGGCAAAAGATAAAACAAGTGTTGCTG |
Secondary |
3D7-A
3D7-B
FC27-A
FC27-B |
GCAGAAAGTAAGCCTTCTACTGGTGCT
GATTTGTTTCGGCATTATTATGA
GCAAATGAAGGTTCTAATACTAATAG
GCTTTGGGTCCTTCTTCAGTTGATTC |
Glurp |
Primary
Secondary |
GLURP-OF GLURP-OR |
TGAATTTGAAGATGTTCACACTGAAC
GTGGAATTGCTTTTTCTTCAACACTAA |
GLURP-OR2
GLURP-NF |
GTGGAATTGCTTTTTCTTCAACACTAA
TGTTCACACTGAACAATTAGATTTAGATCA |
Both primary and secondary PCRs for msp1 and msp2 were carried out in a final volume of 25 μl containing 14.875 μl of sterile water, 0.25 μl of each primer, 2.5 μl of 10X PCR buffer, 2.5 μl of dNTP, 3.5 μl of 25 mM MgCl2, 0.125 μl of One Taq® DNA polymerase and 1 μl of Template DNA or primary PCR product under the following conditions for primary and secondary PCR (30 cycles) : initial denaturation at 95°C for 5 min, extension at 94 °C for 1 min, annealing at 58 °C for 2 min for the primary and 61°C for 2 min for the secondary amplification, extension at 72°C for 2 min, and final elongation at 72°C for 10 min for the primary and 72°C for 5 min for the secondary amplification.
For glurp, the primary and the secondary PCRs were carried out in a total volume of 25 μl containing 12.9 μl of sterile water, 2.5 μl of dNTP, 0.25 μl of each primer, 0.1 μl of One Taq® DNA polymerase, 2.5 μl of 10X PCR buffer and 5 μl of Template DNA or primary PCR product. Temperature cycling parameters (30 cycles) were: initial denaturation at 94°C for 5 min, annealing at 55°C for 2 min, extension at 72°C for 2 min and final extension at 72°C for 10 min. All PCRs were performed in an Applied Biosystems 2720 thermal cycler.
Alleles detection and estimation of molecular weight
The amplified PCR products were either stored at +4°C or analyzed immediately by electrophoresis on a 1.5% agarose gel (Sigma Aldrich Chemie GMBH, Deutschland) on 3% Redsafe® (Nucleic Acid Staining solution). The agarose gels were visualized under UV illumination and then photographed. This allowed a simultaneous typing of the alleles by size polymorphism and identification of the allelic family. The sizes of the PCR products were estimated using 100 bases pairs (bp) DNA ladder marker (Sigma Aldrich Chemie GMBH). The detection of a single PCR fragment for each locus was classified as an infection with one parasite genotype. The detection of more than one PCR fragment for msp1, msp2 or glurp was defined as a multiple P. falciparum infection. The allelic frequency of msp1, msp2 and glurp was calculated as the proportion of the allele detected for each allelic family out of the total of alleles detected. The multiplicity of infection (MOI) was calculated by dividing the total number of alleles detected in msp1, msp2 and glurp by the number of positive samples for each marker. Multiclonality was defined as the percentage of subjects showing more than one genetically distinct parasite type for msp1, msp2 and glurp.
Statistical Analysis
Data were double entered with Excel 2010 and were analyzed using R version 3.5.1 (2018-07-02). The proportions comparisons were made by Chi-squared and normally distributed continuous data by the Student’s t test and ANOVA. The Chi-square test was used to compare proportions. Statistical significance was defined as P < 0.05.
Ethics approval and consent to participate
The study received approval from the Ethics Committee for Health of Burkina Faso before its implementation. It was conducted in accordance with good clinical and laboratory practice. Written informed consent was obtained from each parents or guardians of participant prior to screening and enrolment. Confidentiality of information was ensured by assigning unique identification number to each subject.
Characteristics of the study population
The characteristics of the study population are done in the Table 3. (Table 3). A total of 386 subjects were included. All DNA sample (100%) were successfully amplified for haemoglobin and G6PD. The nested PCR were successfully completed for msp1 and msp2 while 359 (93,00%) were amplified for glurp.
Table 3. Study population characteristics
Parameters |
Value |
IC (95%) |
Mean age (months) |
28.88 |
27.42-30.33 |
Male (n) (%) |
190 (49.22%) |
44.14- 54.32 |
Female (n) (%) |
196 (50.78%) |
45.68-55.86 |
HbAA (n) (%) |
274 (70.98%) |
66.13-75.41 |
HbAC (n) (%) |
94 (24.35%) |
20.22-29.01 |
HbAS (n) (%) |
14 (3.63%) |
2.08- 6.15 |
HbCC (n) (%) |
3 (0.78%) |
0.20-2.45 |
HbSC (n) (%) |
1 (0.26%) |
0.01-1.66 |
G6PD deficiency (n) (%) |
37 (9.59% ) |
6.92-13.08 |
G6PD normal (n) (%) |
349 (90.41%) |
86.92-93.07
|
Parasite density (parasites/µl) |
50,593 |
46,129-55,057 |
n=number of samples
Distribution of msp1, msp2 and glurp alleles
In msp1, the prevalence of K1-type alleles was significantly higher compared to Mad20 and RO33 (p<0.0001). In msp2, the 3D7 allelic family was the most frequently detected (p<0.0001) with 88.08% (340/386) compared to FC27 with 67.36% (260/386). For msp1, twenty-six different alleles were observed according to the band size with fifteen alleles for K1 (fragment range from 120 to 280 bp), ten alleles for Mad20 (fragment range from 120 to 280bp) and one allele for R033 (140bp). In msp2, thirty-five alleles were detected with sixteen alleles for FC27 (fragment range from 160 to 500 bp) and nineteen alleles for 3D7 (fragment range from 120 to 460 bp). For glurp, nine alleles were detected (fragment range from 500 to 950 bp). The prevalence of the three genes msp1, msp2 and glurp are summarized in Table 4. (Table 4).
Table 4. Prevalence of msp1, msp2 and glurp allelic families.
Gene
|
Allele |
Number of P. falciparum isolates |
|
n (%) |
Number of alleles |
Fragment sizes (bp) |
msp1
|
K1 n (%) |
360 (93.26) |
15 |
120-280 |
|
Mad20 n (%) |
305 (79.01%) |
10 |
120-280 |
|
Ro33 n (%) |
324 (83.94%) |
1 |
140 |
p |
<0.0001 |
|
|
msp2 |
3D7n (%) |
340 (88.08%) |
17 |
120-460 |
|
FC27n (%) |
260 (67.36%) |
16 |
160-500 |
p |
<0.0001 |
|
|
|
glurp |
n (%) |
359 (93.00%) |
9 |
500-950 |
n=number of positive samples
Distribution of msp1, msp2 and glurp alleles by genetic factors
None of the three gene (msp1, msp2 and glurp) allelic families were overrepresented in any of the three groups of haemoglobin (Table 5). There were also no statistical differences in the frequencies of msp1, msp2 allelic families and glurp (p=0.70, p=0.93 and p=0.53 respectively) when combining HbC and HbS as abnormal haemoglobin carriers. (Table 5).
Table 5. Prevalence of msp1, msp2 and glurp allelic families by haemoglobin variants
Genes |
Haemoglobin |
Normal haemoglobin
(n=274) |
Abnormal haemoglobin*
(n=112) |
P |
msp1 |
K1 n(%) |
253 (92.34%) |
(n=112) |
0.36 |
Mad20 n(%) |
219 (79.93%) |
86 (76.79%) |
0.58 |
Ro33 n(%) |
231 (84.31%) |
93 (83.04%) |
0.88 |
msp2 |
3D7 n(%) |
244 (89.05%) |
96 (85.71%) |
0.45 |
FC27 n(%) |
180 (65.69%) |
80 (71.43%) |
0.33 |
Glurp n(%) |
258 (94.16%) |
101 (90.18%) |
0.24 |
n=number of positive samples
*Abnormal haemoglobin= HbAC, HbAS, HbCC and HbSC. There is no difference statistically between abnormal haemoglobin types so they have been fused into abnormal haemoglobin
There were no statistical differences (p>0.05) in the frequencies of msp1, msp2 allelic families and glurp by G6PD (Table 6). When combining the three allelic families for msp1 and the two for msp2 there were no differences according to the G6PD status (p = 0.89 and 0.82 respectively). (Table 6).
Table 6. Prevalence of msp1, msp2 and glurp allelic families by G6PD
Genes |
G6PD |
Deficient (n=37) |
Normal (n=349) |
p |
msp1 |
K1 n(%) |
35(94.59%) |
325 (93.12%) |
1.00 |
Mad20 n(%) |
30 (81.08%) |
275 (78.80%) |
0.91 |
Ro33 n(%) |
30 (81.08%) |
294 (84.24%) |
0.79 |
msp2 |
3D7 n(%) |
31 (83.78%) |
309 (88.54%) |
0.56 |
FC27 n(%) |
26 (70.27%) |
234 (67.05%) |
0.83 |
Glurp n (%) |
35(94.59%) |
324 (92.84%) |
0.95 |
n=number of positive samples
Distribution of msp1, msp2 and glurp alleles by fragments sizes
From the msp1 alleles analyzed, three genotypes from K1 (K1180bp, K1200bp and K1220bp); five from Mad20 (Mad20120bp, Mad20160bp, Mad20180bp, Mad20200bp and Mad20220bp) were the most prevalent (Figure 1A-1B). Of the glurp alleles, the glurp700 bp, glurp800bp and glurp900bpwere dominant (Figure 1E). (Figure 1).
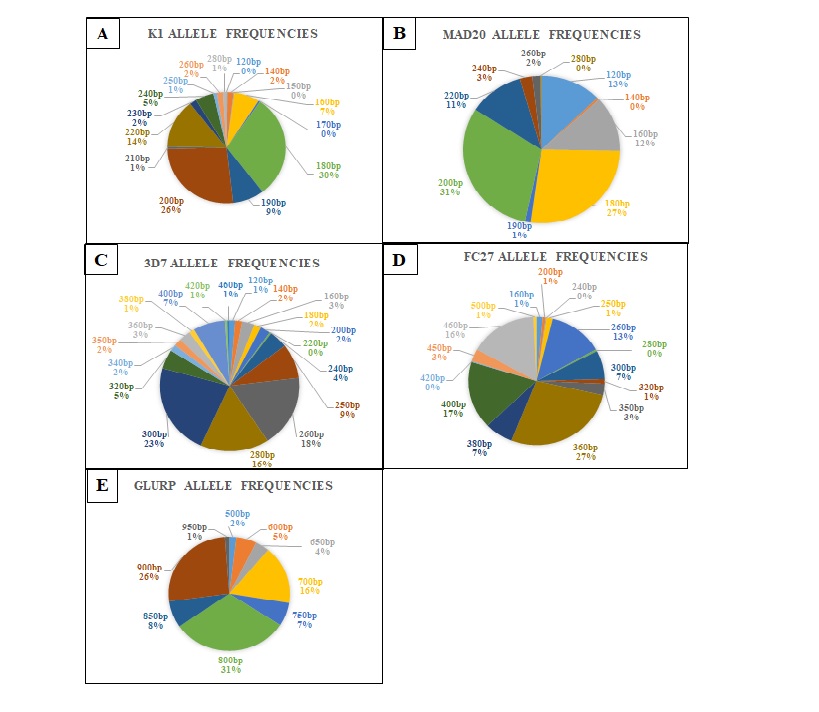
Figure 1. Numbers of P. falciparum msp1 (K1, Mad20), msp2 (3D7 and FC27) and glurp alleles (A-B numbers of msp1 allelic families, C-D numbers of msp2 allelic families and E numbers of glurp allelic families
The prevalence of K1220bp is higher in abnormal haemoglobin than in normal haemoglobin carriers (Figure 2A). The prevalence of the one Ro33 (Ro33140bp) is low in case of abnormality (Figure 2C). (Figure 2).
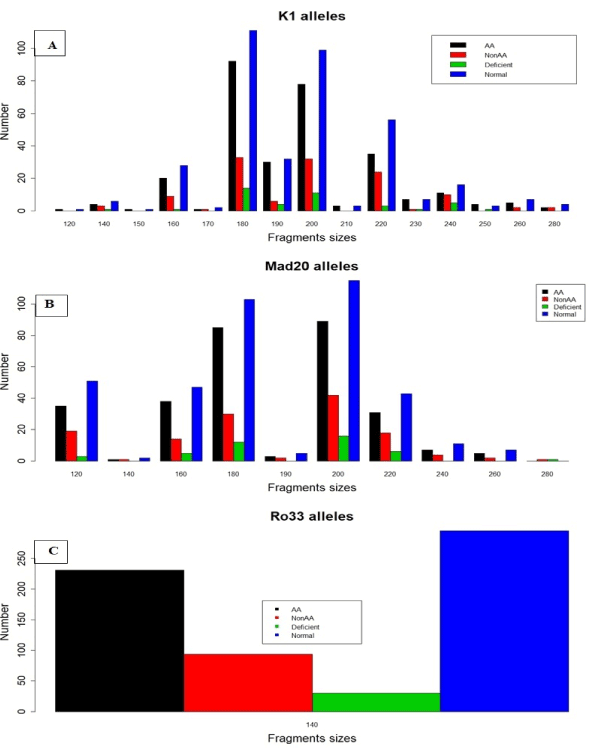
Figure 2. Numbers of P. falciparum msp1 (K1, Mad20 and Ro33) alleles by genetic factors (A-C numbers of msp1 allelic families). AA= Normal haemoglobin, NonAA=Abnormal haemoglobin (HbAC, HbAS, HbCC, HbSC) Deficient= G6PD Deficient and Normal= G6PD Normal`
The prevalence of K1220bp is higher in abnormal haemoglobin than in normal haemoglobin carriers (Figure 2A). The prevalence of the one Ro33 (Ro33140bp) is low in case of abnormality (Figure 2C).
Msp2 3D7250bp, 3D7260bp, 3D7280bp and 3D7300bp were the most prevalent in the study population with statistical difference with 3D7250bp (more prevalent in deficiency cases) and 3D7300bp (more prevalent in normal G6PD carriers) in the G6PD type (p<0.0001 and p=0.001); the FC27360bp, FC27400bp, and FC27460bp were the most prevalent in all the groups (Figure 3A-3B). The glurp700 bp, glurp800bp and glurp900bp (Figure 3C) were dominant in all groups. (Figure 3).
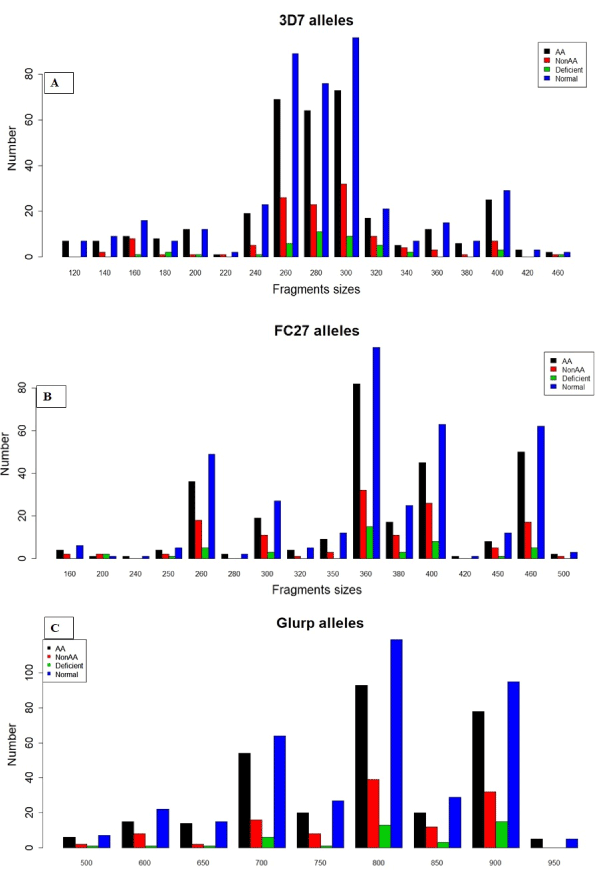
Figure 3. Numbers of P. falciparum msp2 (3D7 and FC27) and glurp alleles by genetic factors (A-B numbers of msp2 allelic families and C numbers of glurp allelic families). AA= Normal haemoglobin, No AA=Abnormal haemoglobin (HbAC, HbAS, HbCC, HbSC) Deficient = G6PD Deficient and Normal= G6PD Normal
MOI and frequency of multiple infections
There was no statistical difference in the MOI of msp1, msp2 and glurp when analyzing the data according to human genetics factors (haemoglobin and G6PD variants) even the mean MOI for msp1 present high values (2.96 and 3.12 for normal and abnormal haemoglobin respectively). (Table 7).
Table 7. Frequency of multiple infections and multiplicity of infection by haemoglobin variants and G6PD
|
Genes |
|
Haemoglobin |
G6PD |
Total |
|
|
|
Normal(274) |
Abnormal(112) |
p |
Normal(349) |
Deficient(37) |
P |
|
|
msp1 |
Frequency n(%) |
255(93.06%) |
103(91.96%) |
0.87 |
324(92.84%) |
34(91.89%) |
1.00 |
358(92.75%) |
|
MOI |
2.96 |
3.12 |
0.19 |
3.00 |
3.08 |
0.67 |
3.01 |
|
msp2 |
Frequency n(%) |
189(68.98%) |
80(71.43%) |
0.72 |
243(69.63%) |
26(70.27%) |
1.00 |
269(69.69%) |
|
MOI |
2.28 |
2.27 |
0.92 |
2.27 |
2.30 |
0.91 |
2.28 |
|
glurp |
Frequency n(%) |
39(14.23%) |
15(13.39%) |
0.92 |
48(13.75%) |
6(16.21%) |
0.87 |
54 (13.99%) |
|
MOI |
1.11 |
1.06 |
0.42 |
1.10 |
1.11 |
0.89 |
1.10 |
Abbreviations: n=number of samples with multiclonal infections for msp1, msp2 or glurp
In the context of malaria immunity affected by both human and different environmental factors, the understanding of the role of human genetic factors in malaria susceptibility still remains an interesting scientific challenge. Thus, many studies far have been concerned with the genetics of host erythrocytes and interestingly they have concerned genes affecting immune processes. Todays, at least two genes affecting red cells confer relative resistance to P. falciparum: the autosomal gene for haemoglobin S (HbS) and the sex-linked gene for the G6PD. The present study was focused on the haemoglobin variants and the mutation A202G of G6PD gene considered to be the most common G6PD deficiency allele in sub-Saharan Africa [15], and analyze their impact on the P. falciparum diversity. It is clearly known that the parasite genotypic and phenotypic diversity of malaria parasites enhances their ability to counteract control measures such as therapeutic drugs.
Consistent with earlier findings, genotyping the two msp markers msp1 and msp2 identified significantly higher numbers of allele variants than genotyping glurp in our study [16]. For msp1, the K1 alleles were more prevalent than the RO33 and Mad 20 alleles. This feature is consistent with previous studies done in Burkina Faso [17] in Benin [18], in Uganda [19], and in Thailand [20]. The number of the msp1 alleles identified is consistent with the msp1 genetic diversity characterized from previous studies carried out in Gabon by Abouy [21] in 2003, by Salem in Mauritania [22] in 2014 and in BF by Somé [17] in 2018. However, the results contrasted with a higher diversity observed in Burkina Faso in 2009 [16]. The different strategies for malaria control in cases management with the wide used of the effective artemisinin-based combination therapies (ACTs) against the resistant genotypes, the wide implementation and coverage of the Impregnated nets, the intermittent preventive treatment (IPTp) in pregnant women, but also the ongoing of the Seasonal Malaria Chemoprevention (SMC) at the country level could justify the reduction of the numbers of the msp1 alleles in BF. The remarkable detection of a single RO33 allele, confirming the reduced polymorphism of this family but also confirm that RO33 is in general in low frequency through different geographic place as in Myanmar where the presence of this type in parasites from the central Myanmar is rare [23]. Overall, fifteen and ten alleles were found in K1 and Mad20 families respectively. The number of alleles is comparable to what has been found in isolates collected in Libreville, but higher than what has been found in isolates from Colombia [24, 25].
For Msp2, more than 150 alleles have already been counted [26]. The 3D7 type was more abundant than the FC27 type, which was similar to what has been reported from Gabon [4], in Thailand [20] and the Thai-Myanmar border [20]. Interestingly, all three sites shared the same most prevalent Msp2 allele (3D7 type, allele size 450–550 bp). Interestingly the thirty-five alleles detected including sixteen alleles for FC27 (fragment range from 160 to 480pb) and nineteen alleles for 3D7 (fragment range from 120 to 460pb) were closed to 25 distinct alleles, including 15 3D7 and 10 FC27 detected in Burkina Faso in 2018 [17]. In contrasts in Uganda, a high number of 49 different alleles have been identified, 38 3D7 and 11 FC27 [19]. For glurp, nine alleles were detected (fragment range from 500 to 950bp). Nevertheless, the number of alleles detected by this marker remains relatively low compared to others study conducted in Burkina Faso, Malawi and Tanzania [19]. As described above some of the strategy such as antimalarial pressure, the use of LLINs, insecticide indoor residual spraying and other factors as demonstrated in previous studies [16] implemented in Burkina since the last decade could explain the reduction of the number of targeted alleles.
A high frequency of the K1220bp and the one Ro33140bp was observed in abnormal haemoglobin carriers. In parallel the study showed, a high frequency of 3D7250bp observed in case of deficiency compared to a high frequency of 3D7300bp allele in normal G6PD carriers. These differences may suggest a specific immune response of the abnormalities (G6PD and haemoglobin) against these alleles. When combining the three allelic families for msp1 there was no difference according to the G6PD status. This probably reflects the fact that G6PD deficiency does not affect the parasite densities rather the clinical outcome of the disease. In a study conducted by Bouyou-Akotet and colleagues [24], they suggest that if an allelic family is predominantly found in an infected population the subjects will develop immunity against these alleles and the presence of less frequent genotypes will more easily induce symptoms. After the combination of the two msp2 allelic families’ there was no difference according to the G6PD status. Since infections complexity is due to the relationship Host-parasite individual reflection, our study showed that there is no effect of haemoglobin type on msp2 families’ alleles even it was described that human genetic factors (abnormal haemoglobin and G6PD deficiency) can inhibit or slow down the parasite development in the host [12]. However, the differences could also be explained by the fact that the totality of the clonal diversity of a P. falciparum infection is not all accessible in a single sample of peripheral blood because of the daily dynamics of sequestration of genotypes in the blood capillaries [27,28].
No effect of abnormal haemoglobin and G6PD deficiency on MOI of msp1, msp2 and glurp were noted in this study. This might suggest that these human factors would appear to have no influence on the diversity of P. falciparum parasites. The innate immunity considered as the first line of human defense and which can help to establish acquired immunity is not yet established in the subjects with abnormalities. In a study conducted in Ghana (children with symptomatic uncomplicated malaria), the authors reported that MOI was lower in HbAS, and HbSS compared to HbAA [29] without difference statistically significant. Previous studies assessing influence of sickle cell gene on the multiplicity of infection (MOI) in asymptomatic malaria in Africa [30] presented reliable results. In addition, in G6PD deficiency group, factors as pre-hospital treatment status, age, transmission intensity, genotyping methods, and immune status can influence the parasite diversity [31]. The high MOI reported in our study contrast with the actions taken to control malaria such as the wide deployment of ACTs, the free distribution campaigns of Long-Lasting Impregnated Mosquito Nets (LLINs) organized since 2010 throughout the country and the ongoing implementation of the SMC. The use of these mosquito nets would seem to have had no impact on the multiplicity of malaria. In all the groups, the polyclonality of msp1, and msp2 values were high. This can be due to factors such as drug pressure, non-compliance with antimalarial prescriptions, self-medication, poor quality antimalarial (forgery, under dosing) that are common in the country. In a study conducted in Colombia, they suggest that it may be the result of a natural process that was halted by the introduction of new anti-malarial treatments [25].
Although the genetic factors (abnormal haemoglobin and G6PD deficiency) studied in our study did not reveal any impact on the genetic diversity of P. falciparum, the existence of other humans genetics factors such as alpha and beta thalassemia, TNFα promoter mutation, HLA factor, membrane traits described by previous studies as conferring a degree of resistance against malaria [8,9] could, however, have an influence on this diversity.
Considering that malaria transmission is endemic and seasonal in Burkina Faso, this study confirms the presence of a large variety of P. falciparum parasites suggesting that there is a need to pursue improving malaria control efforts. K1 and 3D7 are the most prevalent allelic families of msp1 and msp2 respectively. Only one Ro33 allele was observed. In our study no effect of human genetic factors (haemoglobin abnormalities and G6PD deficiency) on the different alleles, the polyclonality and the MOI of msp1, msp2 and glurp genes were observed. In a context of development and design an effective vaccine, such studies will predict the typical behavior of one group over another and improve management of the different strategy.
ESB contributed to the conception and design, acquisition of data, analysis and interpretation of data and wrote the first draft of the manuscript. SBS ABT and ECB designed the study and have final approval of the version to be published. ECB, coordinated the study, collected data coordinated the study, participated in the procedures and revising the manuscript critically for important intellectual content. AO and IS participated in the procedures and revising the manuscript critically for important intellectual content. IN and JBY revising the manuscript critically for important intellectual content. AD SS, SSS, contributed to the acquisition of data. ALO: contributed to data management and data analysis. YT participated in the procedures and revising the manuscript critically for important intellectual content. SBS agreement to act as guarantor of the work. All the authors read and approved the final version.
We thank the population and local authorities of the health district of Saponé and Banfora for their patience and commitment. We are grateful to the staff of the Centre National de Recherche et de Formation sur le Paludisme (CNRFP).
No funding was received for this study.
The authors declare that they have no competing interest.
- WHO. (World Health Organization) World Malaria Report 2018.
- Konaté L, Zwetyenga J, Rogier C, Bishoff E, Fontenille D (1999) Variation of Plasmodium falciparum MSP-1 block2 and MSP-2 allelic prevalence and infection compexity in two neighbouring Senegalese villages with different transmission conditions. Trans R Soc Trop Med Hyg 93: 21-28. [Crossref]
- Doritchamou JYA (2014) Caractérisation des parasites du paludisme gestationnel et optimisation du potentiel vaccinal de VAR2CSA [Thèse en Parasitologie – Biologie cellulaire et moléculaire]: Université Paris Descartes.
- Ntouni F, Ngoundou-Landji J, Luty AJF, Dubreuil G, Millet P (2001) Polymorphisme allélique du gène MSP-2 de Plasmodium falciparum analysé à partir d'échantillons sanguins d'enfants gabonais. Bull Soc Pathol Exot 94: 183-187
- Sondo Paul (2014) Etude de la relation entre le polymorphisme génétique de Plasmodium falciparum et les signes du paludisme à Nanoro, Burkina Faso Université Polytechnique de Bobo-Dioulasso.
- Yazdani SS, Mukherjee P, Chauhan VS, Chitnis CEI (2006) Immune responses to asexual blood-stages of malaria parasites. Curr Mol Med 6: 187-203. [Crossref]
- Mackinnon MJ, Mwangi TW, Snow RW, Marsh K, Williams TN (2005) Heritability of malaria in Africa. PLoS Med 2: e340. [Crossref]
- Modiano D, Luoni G, Sirima BS, Simpore J, Verra F, Konate A, et al. (2001) Haemoglobin C protects against clinical Plasmodium falciparum malaria. Nature 414: 305-308. [Crossref]
- Hill A (1998) The immunogenetics of human infectious diseases. Annu Rev Immunol 16: 593-617. [Crossref]
- DGESS. Direction Générale des Etudes et des Statistiques Sectorielles/Ministère de la Santé, Ouagadougou 2014.
- Plowe CV, Djimde A, Bouare M, Doumbo O, WelllemsTE (1995) Pyrimethamine and proguanil resistance-conferring mutations in Plasmodium falciparum dihydrofolate reductase: polymerase chain reaction methods for surveillance in Africa. Am J Trop Med Hyg 52: 565-568. [Crossref]
- Bwayo D, Kaddumukasa M, Ddungu H, Kironde F (2014) Prevalence of glucose-6-phosphate dehydrogenase deficiency and its association with Plasmodium falciparum infection among children in Iganga distric in Uganda. BMC Res notes 7: 372-376. [Crossref]
- WHO (2007) Methods and techniques for clinical trials on antimalarial drug efficacy: genotyping to identify parasite populations. Amsterdam The Netherlands Geneva: Medicines for Malaria Venture Informal consultation.
- Snounou G, Zhu X, Siripoon N, Jarra W, Thaithong S, Brown KN, et al. (1999) Biased distribution of MSP1 and MSP2 allelic variants in Plasmodium falciparum populations in Thailand. Trans R Soc Trop Med Hyg 93: 369-374. [Crossref]
- Scriver C (1995) Glucose-6-Phosphate Dehydrogenase Deficiency. McGraw Hill: New York.7th edn
- Soulama I, Nébié I, Ouédraogo A, Gansané A, Diarra A, et al. (2009) Plasmodium falciparum genotypes diversity in symptomatic malaria of children living in an urban and a rural setting in Burkina Faso. Malar J 8: 135-142. [Crossref]
- Somé AF, Bazié T, Zongo I, Yerbanga RS, Nikiéma F, et al. (2018) Plasmodium falciparum msp1 and msp2 genetic diversity and allele frequencies in parasites isolated from symptomatic malaria patients in Bobo-Dioulasso, Burkina Faso. Parasit Vectors 11: 323-330. [Crossref]
- Issifou S, Djikou S, Sanni A, Lekoulou F, Ntoumi F (2001) Pas d'influence de la saison de transmission ni de l'âge des patients sur la complexité et la diversité génétique des infections dues à Plasmodium falciparum à Cotonou (Bénin). Bull Soc Pathol Exot 94: 195-198.
- Mwingira F, Nkwengulila G, Schoepflin S, Sumari D, Beck HP, et al. (2011) Plasmodium falciparum msp1, msp2 and glurp allele frequency and diversity in sub-Saharan Africa. Malar J 10: 79-88. [Crossref]
- Congpuong K, Sukaram R, Prompa Y, Dornae A (2014) Genetic diversity of the msp-1, msp-2, and glurp genes of Plasmodium falciparum isolates along the Thai-Myanmar borders. Asian Pac J Trop Biomed 4: 598-602. [Crossref]
- Aubouy A, Migot-Nabias F, Deloron P (2003) Polymorphism in two merozoite surface proteins of Plasmodium falciparum isolates from Gabon. Malar J 2: 12 [Crossref]
- Salem MSA, Ndiaye M, OuldAbdallahi M, Lekweiry KM, Bogreau H, et al. (2014) Polymorphism of the merozoite surface protein-1 block 2 region in Plasmodium falciparum isolates from Mauritania. Malar J 13: 1-8.
- Kang JM, Moon SU, Kim JY, Cho SH, Lin K, et al. (2019) Genetic polymorphism of merozoite surface protein-1 and merozoite surface protein-2 in Plasmodium falciparum field isolates from Myanmar. Malar J 9: 131. [Crossref]
- Bouyou-Akotet MK, M’Bondoukwé NP, Mawili-Mboumba DP (2015) Genetic polymorphism of merozoite surface protein-1 in Plasmodium falciparum isolates from patients with mild to severe malaria in Libreville, Gabon. Parasite 22: 1-9. [Crossref]
- Maestre A, Arango E, Carmona-Fonseca J (2013) Status of allele frequency and diversity of Plasmodium falciparum msp1, msp2 and glurp before implementation of an artemisinin-based combined therapy in Northwestern Colombia. Colomb Med 44: 208-212. [Crossref]
- Silué KD, Felger I, Utzinger J, Beck HP, Smith TA, et al. (2006) Prévalence, diversité antigénique et multiplicité d'infections de Plasmodium falciparum en milieu scolaire au centre de la Côte d'Ivoire. Med Trop 66: 149-156.
- Daubersies P, Sallenave-Sales S, Magne S, Trape JF, Contamin H, et al. (1996) Rapid turnover of Plasmodium falciparum populations in asymptomatic individuals living in a high transmission area. Am J Trop Med Hyg 54:18-26. [Crossref]
- Farnert A, Snounou G, Rooth I, Bjorkman A (1997) Daily dynamics of Plasmodium falciparum subpopulations in asymptomatic children in a holoendemic area. Am J Trop Med Hyg 56: 538-547. [Crossref]
- Mockenhaupt FP, Ehrhardt S, Otchwemah R, Eggelte TA, Anemana SD, et al. (2004) Limited influence of haemoglobin variants on Plasmodium falciparum msp1 and msp2 alleles in symptomatic malaria. Trans R Soc Trop Med Hyg 98: 302-310. [Crossref]
- Ntoumi F, Rogier C, Dieye A, Trape J, Millet P, et al. (1997) Imbalanced distribution of Plasmodium falciparum MSP-1 genotypes related to sickle cell trait. Mol Med. 3: 581-92. [Crossref]
- Patel DK, Mashon RS, Purohit P, Meher S, Dehury S, et al. (2015) Influence of Sickle Cell Gene on the Allelic Diversity at the msp-1 locus of Plasmodium falciparum in Adult Patients with Severe Malaria. Mediterr J Hematol Infect Dis 7: 1-8. [Crossref]



