We previously reported the catalytic activity of 9-mer peptide, JAL-TA9 (YKGSGFRMI), derived from the Box A region of Tob/BTG family of proteins. This was the first report of the catalytic activity of shorter synthetic peptides. Therefore, we used ‘Catalytide’ (catalytic peptide) as the general term for peptides possessing the hydrolase activity. NMR study of JAL-TA9 suggested that the minimum sequence required for the catalytic activity is a 5-mer peptide (GSGFR). In this study, we examined the catalytic activity of this 5-mer peptide against Aβ fragment peptides, Aβ1-20 and Aβ11-29, to find the next Catalytide. Activity of all peptides identified as Catalytides, especially, GQAYR (BTG3) and GQAFR (BTG4), was higher activity than that of GSGFR (Tob1 and 2) and GSGYR (BTG1 and 2). The cleavage mechanism of these Catalytides is still not well understood and needs further investigation. Nonetheless, 5-mer Catalytides are attractive candidates for the development of peptide drugs as a new strategy for treating Alzheimer’s disease (AD).
catalytide, hydrolase activity, Tob/BTG, Box A region, 5-mer synthetic peptide, amyloid-β peptide, alzheimer’s disease
Abbreviations:
APPL: Amyloid Precursor Protein; Aβ: Amyloid Beta; AD: Alzheimer’s Disease; HSA: Human Serum Albumin; TFA: Trifluoroacetic Acid; HPLC: High-Performance Liquid Chromatography; NMR: Nuclear Magnetic Resonance
We recently reported a synthetic nona-peptide, JAL-TA9 (YKGSGFRMI), which possesses proteolytic activity. Our study was the first to report peptide enzyme. Therefore, we used the term Catalytide (Catalytic peptide) for the shorter proteolytic peptides [1]. JAL-TA9 is derived from the Box A region of Tob1, a member of the Tob/BTG family comprising BTG1, BTG2, BTG3/ANA, BTG4, and Tob2. The Box A region of Tob/BTG family is a highly conserved homology domain. According to the recent reports, these proteins display antiproliferative activity in a variety of cell types and are involved in the regulation of tumorigenesis. However, the role of Box A region of Tob/BTG family of proteins remains unclear [2-7].
JAL-TA9 (YKGSGFRMI) is the first Catalytide reported to have auto-proteolytic and hydrolase activities against Aβ fragment peptides. In addition, the secondary structure of JAL-TA9 predicted using NMR, suggested that the 5-mer peptide (GSGFR) might possess the catalytic activity similar to that of JAL-TA9 [1]. In the current study, we evaluated the proteolytic activity of 4 types of 5-mer synthetic peptides derived from the Box A region in Tob/BTG family of proteins (SFig 1) to find a new Catalytides like JAL-TA9 and understand the hydrolase activity of Tob/BTG family proteins.
Chemical peptide synthesis
The peptides were synthesized from fluorenylmethyloxycarbonyl chloride (Fmoc)-protected l-amino acid derivatives according to the method described by Kojima et al. using an automated peptide synthesizer (model 433A, Applied Biosystems, California, USA, 0.1 mmol scale with preloaded resin) [8]. After deprotection according to the manufacturer’s protocol, each peptide was purified using reverse-phase high-performance liquid chromatography (HPLC, Capcell Pak C18 column, SG, 10 or 15 mm i.d. × 250 mm; Shiseido Co., Ltd. Japan) with linear gradient elution from 0.1% trifluoroacetic acid (TFA) to 50% or 70% CH3CN containing 0.1% TFA over 30 min. The flow rate was set to 3.0 or 6.0 mL/min. The primary peak was collected and lyophilized. The purity of the synthetic peptides and the progress of the enzymatic reaction were confirmed using an analytical reverse-phase HPLC (Capcell Pak C18 column, MGII, 4.6 mm i.d. × 150 mm; Shiseido Co., Ltd., Japan), and the flow rate was set to 1.0 mL/min. Linear gradient elution was carried out from 0.1% TFA to 70% CH3CN containing 0.1% TFA. The column eluate was monitored with a photodiode-array detector (SPD-M20A; Shimadzu, Japan). Each purified peptide was characterized by electrospray ionization (ESI)-mass spectrometry (MS) using a Qstar Elite Hybrid LC-MS/MS system [8].
Analysis of proteolytic activity and determination of cleavage sites
5-mer peptides (final concentration of 0.2 mM) were individually incubated with the peptide fragment derived from the Aβ42 (final concentration of 0.05 mM) in the presence of human serum albumin (HSA) (final concentration of 0.025% w/v) in phosphate buffered saline (PBS, pH 7.4) at 37 °C. Ten microliters of each reaction mixture was analyzed in a time-dependent manner using the analytical HPLC system described above. For peak collection, 20 μL of the reaction mixture was loaded into the same HPLC system, and the peak fractions monitored at 220 nm were collected into microtubes (Eppendorf Safe-Lock Tubes, 1.5 mL).
After lyophilization, an appropriate quantity, as determined from the chromatographic peak heights, of 36% CH3CN containing 0.1% HCOOH was added with stirring by an automatic mixer. The cleavage sites were determined by ESI-MS using the flow injection method with 70% CH3CN containing 0.1% HCOOH on a Qstar Hybrid LC-MS/MS system (ABI), and the flow rate was set to 0.1 mL/min [12].
We synthesized 4 types of 5-mer synthetic peptides, GSGFR, GSGYR, GQAYR, and GQAFR which were derived from the Box A region of Tob/BTG family proteins (SFig. 1). Each peptide was purified using a reversed-phase preparative HPLC and the purity of each peptide was checked using an analytical HPLC system using a photo-diode array detector. The molecular weights of purified peptides were confirmed with MS (SFig. 2). On the basis of these data, we judged that all peptides were of adequate purity for carrying out the following experiments.
The proteolytic activity was analyzed as reported previously [1]. Each peptide (final concentration 0.2 mM) was co-incubated with Aβ1-20 or Aβ11-29 as substrates (final concentration 0.05 mM) in PBS at 37 °C in the presence of HSA (human serum albumin), and 10 μL of the reaction mixture was subsequently analyzed by an analytical HPLC on day 0, 2, 4, and 5 (SFig. 3 and 4). Aβ1-20 and Aβ11-29, fragment peptides of Aβ peptide, were stable in PBS containing HSA (SFig 3). The proteolytic activity was evaluated by the decrease in substrate concentration. For GSGFR and GSGYR, the peak heights of Aβ1-20 and Aβ11-29 slightly decreased; however, no new peak was observed on the chromatogram even in day 2 sample, indicating that the proteolytic activities of GSGFR and GSGYR were very weak (Figures 1a & 1b). On the other hand, GQAYR and GQAFR completely cleaved Aβ1-20 in 2 days, and several new peaks were observed (Figures 1c & 1d).
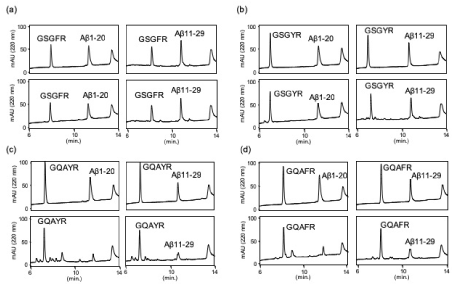
Figure 1. Chromatogram of reaction mixtures. (a) GSGFR, (b) GSGYR, (c) GQAYR or (d) GQAFR were co-incubated with (left) Aβ1-20 or (right) Aβ11-29 in PBS containing HSA, and 10 μL of the reaction mixture was analyzed at 0 (upper) and 2 days (lower)
The decreasing ratio of 4 peptides against Aβ1-20 and Aβ11-29 is shown in Figure 2. These data indicate that the proteolytic activities of GQAYR and GQAFR against Aβ1-20 and Aβ11-29 are stronger than those of GSGFR and GSGYR.
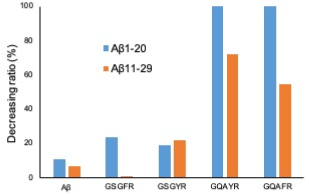
Figure 2. Decreasing ratio of each substrate co-incubate with 5-mer peptide. The decreasing ratio was calculated by the peak height obtained on day 0 and day 2 in HPLC analysis
We next identified the cleavage sites on Aβ1-20 and Aβ11-29 when using GQAYR or GQAFR. The peak heights of both 5-mer peptides and substrates decreased in a time dependent manner. When Aβ1-20 was used as a substrate, it almost disappeared on day 2; however, GQAYR and GQAFR were still remained. Therefore, we continued the incubation (SFig. 4) and collected every peak that appeared on day 5 by monitoring at 220 nm (Figure 3). Additional 12 (A1-10) and 4 peptides (C1-4) were identified as fragment peptides derived from Aβ1-20 and GQAYR, respectively (Figure 3a). For GQAFR, 12 (A1-7) and 5 peptides (C1-5) were also identified as fragment peptides derived from Aβ1-20 and GQAFR, respectively (Figure 3b). The cleavage sites on Aβ1-20 corresponding to GQAYR and GQAFR are shown in Fig. 3c. These data indicate that GQAYR and GQAFR possess the auto-proteolytic activity as well as can cleave the Aβ1-20 similar to JAL-TA9 [1].
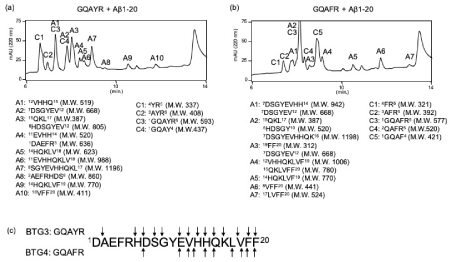
Figure 3. Determination of cleavage site against Aβ1-20. On day 5, 20 μL of reaction mixture was injected and all peaks were collected. Aβ1-20 was co-incubated with (a) GQAYR or (b) GQAFR. (c) Cleavage sites
We next identified the cleavage sites on Aβ11-29. Eight (A1-7) and four peptides (C1-4) were identified as fragment peptides derived from Aβ11-29 and GQAYR, respectively (Figure 4a). For GQAFR, 12 (A1-7) and 4 peptides (C1-4) were also identified as fragment peptides derived from Aβ11-29 and GQAFR, respectively (Figure 4b). The cleavage sites on Aβ11-29 corresponding to GQAYR and GQAFR are shown in Figure 4c. Interestingly, no fragment peak derived from HSA was identified in any of the above cases. The other peptides GSGFR and GSGYR, also showed the proteolytic activity (SFig. 5).
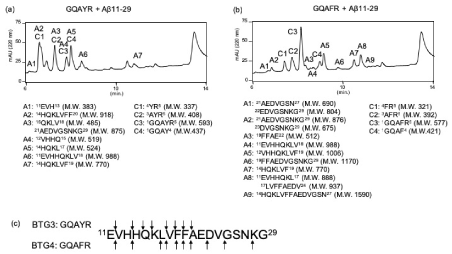
Figure 4. Determination of cleavage site against Aβ11-29. On day 5, 20 μL of reaction mixture was injected and all peaks were collected. Aβ1-20 was co-incubated with (a) GQAYR or (b) GQAFR. (c) Cleavage sites
Alzheimer’s disease (AD) is the most common age-related neurodegenerative disorder caused by the aggregation and accumulation of Aβ42. Many trials have been conducted to develop drugs for the treatment of AD, however, the results have not been encouraging [9-14]. Thus, new effective drugs are urgently required for treating AD. Among 4 peptides tested in this study, GQAYR and GQAFR strongly cleaved Aβ fragment peptides, especially Aβ1-20, with many cleavage sites, (Figures 3 & 4) suggesting that these 5-mer peptides can cleave Aβ42.
The antiproliferative ability of the Tob/BTG family of proteins is due to their association with the target protein in the cell. The highly conserved region in N-terminal region of the Tob/BTG, Box A, mediates the majority of the Tob/BTG interactions. However, no study has reported the hydrolase activity of Tob/BTG family proteins themselves or their fragment peptides. In this study, we found the proteolytic activity of the synthetic 5-mer peptides derived from the Box A region of Tob/BTG family proteins; especially GQAYR from BTG3 and GQAFR from BTG4 fragment peptide showed potent activity (Figure 2) with many cleavage sites (Figures 3 & 4). These results suggest that Tob/BTG family proteins may also possess previously unknown proteolytic activity.
The mechanism underlying the proteolytic activity remains unexplored. To elucidate this mechanism, stereo-structure analyses are required. Therefore, we attempted to predict the secondary structure in silico on a personal computer using MM2 and MMFF94 programs. The stereo-structures of GSGFR and GSGYR were similar to each other. In addition, the location of three amino acids, Arg-Phe/Tyr-Ser, seemed to be close (SFig. 6a and b). On the other hand, the stereo-structure and the location of Arg, Phe/Tyr-Gln were also similar to each other but were different from those in GSGFR and GSGYR (SFig. 6c and d). These results may relate to the proteolytic activity (Figure 2). We are currently investing the stereo-structures of 5-mer peptides using NMR analysis.
We conclude that 5-mer peptides derived from the Box A region of Tob/BTG family of proteins cleave Aβ fragment peptides and are categorized as Catalytides. Therefore, GQAYR and GQAFR can also be categorized as the Catalytides. Along with JAL-TA9, these Catalytides can prove as attractive drug candidates for the prevention and treatment of AD.
Availability of data and material
All data generated or analysed during this study are included in this published article.
The authors declare that they have no competing interests.
T. A. and R. N. were responsible for the experimental design and data interpretation. T. A., R. N., Y. H., and M. S. mainly contributed to writing and reviewing the manuscript. R. N. and M. K. conducted all of the experiments, especially the HPLC analyses and the determination of cleavage sites. R. N. and M. K. contributed to the MS analyses. M. K., and R. N. contributed to the structure analyses using NMR and the computer modeling.
This work was partially supported by the Japan Society for the Promotion of Science (JSPS) Grants-in-Aid for Scientific Research (KAKENHI) Program, Grant Number 15K07908, and the Okinawa Institute of Science and Technology (OIST) Proof-of-Concept (POC) Program.
View supplementary data
- Nakamura R, Konishi M, Taniguchi M, Hatakawa Y, Akizawa T (2019) The discovery of shorter synthetic proteolytic peptides derived from Tob1 protein. Peptides 116: 71-77.
- Winkler GS (2010) The mammalian anti-proliferative BTG/Tob protein family. J Cell Physiol 222: 66-72. [Crossref]
- Schulze-Topphoff U, Casazza S, Varrin-Doyer M, Pekarek K, Sobel RA, et al. (2013) Tob1 plays a critical role in the activation of encephalitogenic T cells in CNS autoimmunity. J Exp Med 210: 1301-1309. [Crossref]
- Chen Y, Wang C, Wu J, Li L (2015) BTG/Tob family members Tob1 and Tob2 inhibit proliferation of mouse embryonic stem cells via Id3 mRNA degradation. Biochem Biophys Res Commun 462: 208-214.
- Yang X, Morita M, Wang H, Suzuki T, Yang W, et al. (2008) Crystal structures of human BTG2 and mouse TIS21 involved in suppression of CAF1 deadenylase activity. Nucleic Acids Res 36: 6872-6881.
- Matsuda S, Kawamura-Tsuzuku J, Ohsugi M, Yoshida M, Emi M, et al. (1996) Tob, a novel protein that interacts with p185erbB2, is associated with anti-proliferative activity. Oncogene.
- Horiuchi M, Takeuchi K, Noda N, Muroya N, Suzuki T, et al. (2009) Structural basis for the antiproliferative activity of the Tob-hCaf1 complex, J Biol Chem 284: 13244-13255.
- Kojima S, Konishi M, Akizawa T (2014) Prion fragment peptides are digested with membrane type matrix metalloproteinases and acquire enzyme resistance through Cu2+-binding. Biomolecules 4: 510-526.
- Doody RS, Thomas RG, Farlow M, Iwatsubo T, Vellas B, et al. (2014) Phase 3 Trials of Solanezumab for Mild-to-Moderate Alzheimer’s Disease. N Engl J Med.
- T Sato, Kienlen-Campard P, Ahmed M, Liu W, Li H, et al. (2006) Inhibitors of amyloid toxicity based on ß-sheet packing of Aß40 and Aß42. Biochemistry.
- Kumar J, Namsechi R, Sim VL (2015) Structure-based peptide design to modulate amyloid beta aggregation and reduce cytotoxicity. PLoS One 10: 1-18.
- Young LM, Saunders JC, Mahood RA, Revill CH, Foster RJ, et al. (2015) Screening and classifying small-molecule inhibitors of amyloid formation using ion mobility spectrometry-mass spectrometry. Nat Chem.
- Crouch PJ, Tew DJ, Du T, Nguyen DN, Caragounis A, et al. (2009) Restored degradation of the Alzheimer’s amyloid-ß peptide by targeting amyloid formation. J Neurochem 108: 1198-1207.
- Storr T, Merkel M, Song-Zhao GX, Scott LE, Green DE, et al. (2007) Thompson, B.O. Patrick, H.J. Schugar, C. Orvig, Synthesis, characterization, and metal coordinating ability of multifunctional carbohydrate-containing compounds for Alzheimer’s therapy. J Am Chem Soc 129: 7453-7463.




