Gastric cancer is one of the most common malignancies and remains the third leading cause of cancer-related death worldwide. We previously developed a method of DNA methylation analysis for early detection of gastric cancer in gastric washes rather than the highly acidic gastric juice. Exosomes in gastric juice may provide an alternative to gastric washes for the molecular detection of gastric cancer. We sought to determine whether exosomes can be purified from the gastric juice of gastric cancer patients and whether the exosomes contain significant amounts of tumor-related methylated DNA. Using a polymer-based reagent, we purified exosomes from the culture media of gastric cancer cell lines as well as gastric juice of patients with gastric cancer. Methylation levels of LINE1 and tumor-related SOX17 gene in exosomal DNA were analyzed by bisulfite pyrosequencing. The microvesicles were verified as exosomes by transmission electron microscopy and western blotting with the CD9 exosomal marker. LINE1 methylation were reduced in nuclear DNA and in the corresponding exosomal DNA in gastric cancer cell lines. Concordant methylation levels of SOX17 gene were observed in exosomal and nuclear DNA in gastric cancer cell lines, suggesting the methylated DNA is efficiently packaged in exosomes. Using exosomal DNA derived from gastric juice, we were also able to detect SOX17 DNA methylation, which reflects the nuclear DNA methylation status of the corresponding tumor. SOX17 methylation was detected in both early and advanced gastric cancer of intestinal and diffuse types. These findings expand the functional molecular content of tumor exosomes to include tumor-related methylated DNA and suggest a potential use for methylation analysis of exosomal DNA derived from gastric juice as a biomarker for gastric cancer.
DNA methylation, exosome, gastric cancer gastric juice, SOX17
Gastric cancer (GC) is the third highest cause of global cancer mortality [1]. GC is a heterogeneous disease with multiple environmental etiologies and alternative carcinogenic pathways [2-6]. The development of noninvasive biomarkers to detect early cancer and/or reflect an individual’s cancer risk is essential to reducing GC mortality [7,8].
Molecular markers in the gastric juice may provide a noninvasive approach to detecting GC. Indeed, gastric juice microRNAs (miR-421, miR-129, miR-21, and miR-106a) and long noncoding RNA (AA174084) have been reported as potential biomarkers for GC screening [9-12]. However, the various cellular sources of miRNAs in gastric juice could make quantitative expression analysis particularly difficult for the molecular detection of GC. Indeed, the use of gastric juice DNA for molecular diagnostics has been deemed unfeasible because the DNA is easily degraded by gastric acidity [13]. One alternative to gastric juice is the use of gastric washes, for which we have developed a method for early GC detection by DNA methylation analysis of genes such as MINT25 and sex determining region Y-Box 17 (SOX17) [13-15].
Another alternative is exosomes, which have received increasing attention [16-19]. Exosomes are small membrane vesicles secreted by various cell types, including cancer cells. Exosomes plays a role in intercellular communication as a delivery system for cells, tissues, and organs. Exosomes carry functional biomolecules such as nucleotides and proteins; they have been detected in a variety of body fluids and in cell culture media in vitro [20]. Because exosomes are very stable under diverse conditions in the extracellular environment, functional biomolecules are protected against degradation and denaturation [21,22]. Therefore, the use of exosomes from gastric juice may provide an alternative to gastric washes for molecular detection of GC.
Because tumor-derived exosomes carry various tumor-related functional biomolecules, they are considered promising candidate diagnostic biomarkers of cancer [16,22,23]. Among them, mRNA, microRNA (miRNA), and protein have been studied extensively. Exosomes also carry retrotransposonRNA transcripts such as LINE1 and Alu elements [24], single-stranded DNA [24], mitochondrial DNA [25,26], and amplified oncogene sequences such as c-myc[24]. Moreover, exosomes from the serum of patients with pancreatic cancer reportedly carry genomic double-stranded DNAs (dsDNAs), which span all chromosomes and contain DNA with mutated KRAS and p53 [22]. Similarly, exosomal DNA (exoDNA) reportedly carries the entire genome and thus reflects the mutational status of the parental cancer cells [27]. Thus, the translational value of exoDNA in tumor-derived exosomes has great potential utility as a biomarker for early detection of cancer and metastasis [27].
Given the frequencies of and detection methods for genetic and epigenetic alterations [6,7], DNA methylation is more appropriate than mutations for molecular detection of GC. Although the overall level of exoDNA 5′-cytosine methylation is similar to that of genomic DNA [27], it is unclear whether exosomes contain tumor-related methylated DNA. In this study, we demonstrated the feasibility of purifying exosomes from the gastric juices of patients with GC; we also sought to determine whether exosomes contain significant amounts of methylated DNA and whether they reflect the DNA methylation status of the corresponding cancer. LINE1 methylation levels as measured by pyrosequencing are good indicators of the cellular levels of 5-methylcytosine (i.e., the global DNA methylation level) [28]. LINE1 methylation was also analyzed by bisulfite pyrosequencing as surrogates for genome-wide hypomethylation.
Cell lines and clinical samples
Nine GC cell lines (AZ521, MKN1, MKN7, MKN45, MKN74, NUGC2, NUGC3, NUGC4, and KatoIII) were obtained from the American Type Culture Collection (Manassas, VA, USA) and the Japanese Collection of Research Bioresources (Tokyo, Japan). All cell lines were maintained in appropriate culture media containing 10% fetal bovine serum (FBS) or exosome-depleted FBS media supplement (System Biosciences (SBI), Mountain View, CA) in plastic tissue culture plates. Gastricjuice samples were obtained from 20 patients with GC (10 early and 10 advanced cancers) before endoscopic treatment or surgery. This study was approved by the Institutional Review Board and informed consent was obtained from each subject.
Isolation of exosomes from culture media and gastric juice
Exosomes were extracted from culture media and gastric juice using ExoQuick-TC Exosome Precipitation Solution (SBI, CA). Samples were centrifuged at 3000×g for 15 min to remove cells and cellular fragments. ExoQuick-TC was added to the supernatants and exosomes were precipitated by refrigeration at -20°C for 12 h. Exosome pellets collected by centrifugation at 1500×g for 30 min were dissolved in 20 µl phosphate-buffered saline (PBS). Exosomes were quantified by using the micro BCA protein assay (Thermo Fisher Scientific KK, Tokyo, Japan).
Western blot analysis
Western blotting was performed as described [14] with primary and secondary antibodies coupled to horseradish peroxydase, diluted according to supplier recommendations. Primary antibodies included anti-CD9 (SBI), anti-calnexin (endoplasmic reticulum marker; ab10286, Abcam, Cambridge, England), anti-GM130 (Golgi apparatus marker; clone 35, BD Biosciences, San Jose, CA), and anti-aconitase 2 (ACO2, mitochondrial marker; HPA, Sigma-Aldrich, St. Louis, MO) [29-31].
Transmission electron microscopy (TEM)
Analysis was performed as described [32]. Aliquots of re-suspended vesicles (5 µl) were placed onto support grids (Cu 200M), and allowed to adsorb for 60 s. Grids were washed twice in double-distilled water, and vesicles were stained for 10 s in a 2% aqueous solution of uranyl acetate. Vesicle size and morphology were examined by TEM (JEM-1200EX, (JEOL, Akishima, Japan).
DNA preparation from exosomes, cell lines, and GC tissues
Formalin-fixed paraffin-embedded (FFPE) specimens were sliced in 10 µm sections and subjected to laser-capture microdissection (LCM) to isolate cancer and normal cells using the PALM Microbeam (Carl Zeiss, Oberkohen, Germany) [33]. DNA was extracted from exosomes and microdissected FFPE tissues by using a standard phenol–chloroform method. Isolated exosomes were treated with DNase before DNA extraction to reduce the chance of external DNA contamination [24]. After exosome lysis and DNA purification, exoDNA was treated with RNase [24]. The concentration and quantity of all DNA extracts were measured by NanoDrop spectrophotometry (ND-1000 Spectrophotometer; NanoDrop Technologies, Wilmington, DE).
LINE1 methylation analysis
To quantify LINE1 methylation levels, we used pyrosequencing technology [28]. PCR and pyrosequencing for LINE1 were performed using the PyroMark kit (Qiagen). This assay amplifies a region of the LINE1 element that includes CpG sites. Cycling conditions were as follows: 45 cycles at 95°C for 20 s, 50°C for 20 s, and 72°C for 20 s, followed by 72°C for 5 min. The biotinylated PCR product was purified and converted to single strands to serve as a template for the pyrosequencing reaction using the Pyrosequencing Vacuum Prep Tool (Qiagen). The pyrosequencing reactions were performed using the PyroMark Q24 (Qiagen). The percentage of Cs relative to the total sum of Cs and Ts at each CpG site was calculated. The average of the percentages of Cs at the CpG sites was used to represent the overall LINE1 methylation levels in each sample.
Bisulfite-pyrosequencing of tumor-related genes
Bisulfite treatment, PCR, and pyrosequencing were performed as described [13]. All primers and PCR conditions for amplifying CpG island DNA fragments of SOX17 were described previously [14].
RNA extraction and quantitative RT-PCR
Total RNA was extracted from microdissected FFPE tissues using the miRNeasy kit (Qiagen). Real-time quantitative RT-PCR of SOX17 was performed with a TaqMan Gene Expression Assay (Applied Biosystems) as described [14].
Statistical analysis
Methylation levels (percentage) were analyzed as a continuous variable for comparison. Spearman’s rank correlation coefficient was used to assess the correlation of DNA methylation between exoDNA and cellular or tissue nuclear DNA. P<0.05 was considered significant.
Identification and characterization of exosomes from culture media of GC cell lines
To isolate exosomes, we used ExoQuick-TC precipitation solution, a polymer-based reagent. To ensure the efficacy of the exosome isolation method, we characterized the microvesicles by western blot analysis. Western blot signal intensities for exosomal marker CD9 were detected in the exosome fraction but not in the cell lysates (Figure 1). In contrast, mitochondrial (ACO2), endoplasmic reticulum (calnexin), and Golgi apparatus (GM130) markers were detected in cell lysates but not in exosomes, indicating efficient enrichment of exosomes from the culture media.
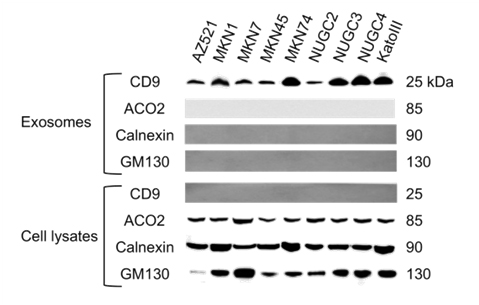
Figure 1. Characterization of exosomes isolated from GC cell lines by western blot analysis. Exosome preparations were positive for the exosomal marker CD9 and negative for proteins from the mitochondria (ACO2), endoplasmic reticulum (calnexin), and Golgi apparatus (GM130), which were detected in cell lysates.
Characterization of exosomesisolated from gastric juice by TEM and western blot analysis
Exosomes were purified from gastric juice samples using the ExoQuick-TC exosome purification kit. We characterized the microvesicles by TEM and western blot analysis. Round particles with a characteristic exosomal size (30–100 nm) and shape were observed (Figure 2a). CD9 expression was specifically observed in isolated exosomes (Figure 2b).

Figure 2. Characterization of gastric juice-derived exosomes by TEM and western blot analysis. (a) Morphorogical TEM characterization of exosomes derived from the gastric juice of patients with GC. Exosomes were purified from gastric juice with the ExoQuick-TC exosome purification kit. Round particles with a characteristic exosomal size (30–100 nm) and shape were observed. (b) Different protein concentrations (10 and 30 µ
µg) of purified exosome lysates were separated by polyacrylamide gel electrophoresis, transferred to membranes, and blotted with anti-CD9 antibody. CD9 expression was specifically observed in isolated exosomes.
Detection of long fragments of DNA in exosomes
After exoDNA extraction from GC cell lines, the samples were subjected to RNase A treatment to exclude RNA, then separated by 2% agarose gel electrophoresis (Figure 3). Long DNA fragments were observed in each sample. Exosomes from GC cell lines contained genomic dsDNA, detected by a double-stranded DNA detection kit (data not shown).
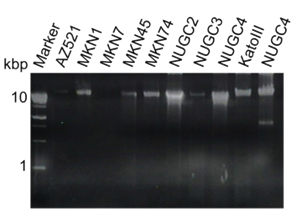
Figure 3. Detection of long fragments of DNA in exosomes from GC cell lines. After exosome lysis and DNA extraction, DNA was treated with RNase A. The samples were separated by 2% agarose gel electrophoresis. Exosomes contained >10-kb fragments of genomic DNA. Cellular genomic DNA of NUGC4 is shown at the rightmost column.
Concordant methylation levels between exosomal and nuclear DNA in GC cell lines
LINE1 methylation levels were similarly low in the nuclear DNA (41.0 ± 10.8) and exoDNA (43.1 ± 10.4) of GC cell lines (Figure 4a, data not shown). Methylation levels were concordant between exosomal and nuclear DNA in GC cell lines (r=0.88, P=0.013). Methylation levels of the SOX17 gene were analyzed by quantitative bisulfite pyrosequencing of exosomal and nuclear DNA in GC cell lines and shown to be concordant (Figures 4b-4d).
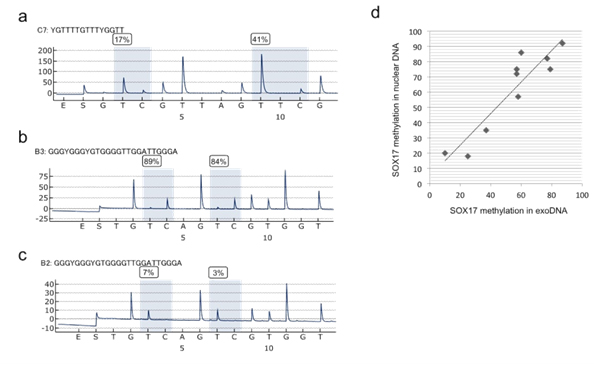
Figure 4. Methylation levels of LINE1 and SOX17 as shown by bisulfite pyrosequencing of exoDNA derived from the culture media of GC cell lines. (a) Pyrogram of LINE1(b) and(c) Pyrogram of the SOX17 gene. (d) Correlation of SOX17 methylation levels between exosomal and nuclear DNA in the same cell line.
Concordant methylation levels between exosomal and tissue nuclear DNA in patients with GC
LINE1 methylation levels were low in nuclear DNA (42.6 ± 10.8) and corresponding gastric juice-derived exoDNA (47.3 ± 11.4) (Figure 5a, data not shown), but were generally lower in the nuclear DNA. Methylation levels were concordant between exosomal and tissue nuclear DNA in patients with GC (r=0.82, P=0.014). Quantitative bisulfite pyrosequencing analysis of SOX17 showed varying levels of methylation in gastric juice-derived exoDNA (Figures 5b-5c and data not shown). Concordant SOX17 methylation levels were observed between exosomal and tissue nuclear DNA from 10 patients with GC (Figure 5d). We further analyzed 10 GC cases and significant SOX17 methylation (>20%) was detected in 16 of 20 GC cases (8 of 10 early GCs and 8 of 10 advanced GCs), but in only one of the 10 control samples without GC. Finally, SOX17 mRNA expression was frequently reduced in GC tissues with SOX17 methylation but not in those without methylation (Figure 5e, data not shown).
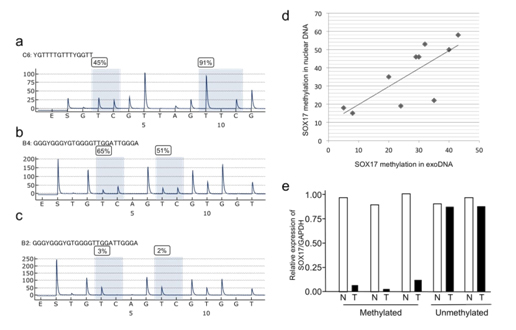
Figure 5. Methylation levels of LINE1 and SOX17 as shown by bisulfite pyrosequencing of exoDNA derived from the gastric juice of patients with GC. (a) Pyrogram of LINE1 (b) and (c) Pyrogram of the SOX17 gene. (d) Correlation of SOX17 methylation levels between gastric juice exoDNA and nuclear DNA from FFPE samples in the same patient. (e) Expression of SOX17 mRNA in GC with or without DNA methylation. Real-time PCR was performed with cDNA from extracted DNA by LCM. N, Nontumor tissue, T, tumor tissues.
This is the first report of exosome purification from the gastric juices of patients with GC. We used a polymer-based reagent and this method has been verified in previous studies [30]. Validation of gastric juice-derived exosomes was performed by TEM and western blotting with an exosomal marker. Characteristic exosome size and shape was observed by TEM; western blotting showed that exosomes expressed the exosomal marker CD9 protein [29], thus demonstrating efficient enrichment of exosomes from gastric juice.
GC-derived exosomes may contain DNA. To test this hypothesis, we extracted DNA from exosomes derived from the culture media of GC cell lines. We showed that exosomes contain genomic dsDNA. Our results also support the notion that exosomes can carry large fragments (>10 kb) of genomic DNA [22,27].
We previously developed a method for GC detection by DNA methylation analysis of gastric washes [13,14]. Therefore, we were interested in using exoDNA for methylation analysis, because exoDNA is not easily damaged by gastric acidity. Indeed, quantitative methylation analysis of LINE1 was feasible and showed that methylation levels were low in nuclear DNA and the corresponding exoDNA in GC cell lines. In general, concordant methylation was observed in exosomal and nuclear DNA of GC cell lines, suggesting that exoDNA reflects the nuclear DNA methylation status of the corresponding tumor.
We next analyzed whether exosomes contain significant amounts of tumor-related methylated DNA that reflects the methylation status of nuclear DNA. To test this hypothesis, methylation levels of the SOX17 gene were analyzed by quantitative bisulfite pyrosequencing [14]. The methylation levels of exosomal and nuclear DNA in GC cell lines were generally concordant, suggesting that tumor-related methylated DNA is packaged in exosomes.
Based on our observations in cell lines, we speculated that gastric juice-derived exosomes may also contain tumor-related methylated DNA. Quantitative methylation analysis of LINE1 revealed low methylation levels in nuclear DNA and corresponding gastric juice-derived exoDNA. Methylation levels were slightly higher in gastric juice-derived exoDNA than in the corresponding nuclear DNA, suggesting contamination of exoDNA derived from non-cancer cells in gastric juice. Thus, quantitative methylation analysis of LINE1 may not provide appropriate biomarker-based detection of GC.
We then analyzed DNA methylation of the SOX17 gene in gastric juice-derived exoDNA samples from patients with GC. SOX17 methylation was detected even in early GC, regardless of differentiation type. Moreover, concordant SOX17 methylation levels were observed between exosomal and tissue nuclear DNA. Selective packaging of retrotransposon RNA sequences, especially HERV, in tumor microvesicles has been reported [24]. The underlying mechanisms of methylated DNA packaging into tumor exosomes need to be further characterized [27].
Not only the clinical utility but also the pathobiological effects of nucleic acids in circulation (nucleosomes, DNA, RNA, microRNA etc.) are receiving increasing attention [34,35]. Further analysis is necessary to clarify the possible effects of shedded epigenetic materials through exosomes in the tumor microenvironment [27].
To our knowledge, this is the first report to demonstrate that gastric juice-derived exoDNA samples reflect the nuclear DNA methylation status of the corresponding tumor. Therefore, gastric juice-derived exoDNA methylation analysis may be useful for detecting GC. Because gastrointestinal endoscopy is costly and painful for patients, it is difficult to incorporate the technique into routine clinical settings, especially in developing countries [11]. In contrast, gastric juice samples can be easily and repeatedly obtained and exoDNA is not easily denatured by gastric acidity [13]. Although our data require further validation, we suggest detection of methylated DNA in gastric juice-derived exosomes may be a novel noninvasive tool for detecting GC.
The authors declare no competing interests.
- 1. Ferlay J (2013) GLOBOCAN 2012 v1.0, Cancer Incidence and Mortality Worldwide: IARC CancerBase No. 11. International Agency for Research on Cancer.
- 2. Ushijima T, Sasako M (2004) Focus 2021 Copyright OAT. All rights reservi>: 121-125. [Crossref]
- 3. Wadhwa R, Song S, Lee JS, Yao Y, Wei Q, et al. (2013) Gastric cancer-molecular and clinical dimensions. Nat Rev Clin Oncol 10: 643-655. [Crossref]
- 4. Yamamoto H, Watanabe Y, Maehata T, Morita R, Yoshida Y, et al. (2014) An updated review of gastric cancer in the next-generation sequencing era: insights from bench to bedside and vice versa. World J Gastroenterol 20: 3927-3937. [Crossref]
- 5. McLean MH, El-Omar EM (2014) Genetics of gastric cancer. Nat Rev Gastroenterol Hepatol.
- 6. Ushijima T, Asada K (2010) Aberrant DNA methylation in contrast with mutations. Cancer Sci 101: 300-305. [Crossref]
- 7. The Cancer Genome Atlas Research Network (2014) Comprehensive molecular characterization of gastric adenocarcinoma. Nature: 202-209. [Crossref]
- 8. Sukawa Y, Yamamoto H, Nosho K, Ito M, Igarashi H, et al. (2014) HER2 expression and PI3K-Akt pathway alterations in gastric cancer. Digestion 89: 12-17. [Crossref]
- 9. Zhang X, Cui L, Ye G, Zheng T, Song H, et al. (2012) Gastric juice microRNA-421 is a new biomarker for screening gastric cancer. Tumour Biol 33: 2349-2355.[Crossref]
- 10. Yu X, Luo L, Wu Y, Yu X, Liu Y, et al. (2013) Gastric juice miR-129 as a potential biomarker for screening gastric cancer. Med Oncol 30: 365. [Crossref]
- 11. Cui L, Zhang X, Ye G, Zheng T, Song H, et al. (2013) Gastric juice MicroRNAs as potential biomarkers for the screening of gastric cancer. Cancer 119: 1618-1626. [Crossref]
- 12. Shao Y, Ye M, Jiang X, Sun W, Ding X, et al. (2014) Gastric juice long noncoding RNA used as a tumor marker for screening gastric cancer. Cancer. [Crossref]
- 13. Watanabe Y, Kim HS, Castoro RJ, Chung W, Estecio MR, et al. (2009) Sensitive and specific detection of early gastric cancer with DNA methylation analysis of gastric washes. Gastroenterology136: 2149-2158. [Crossref]
- 14. Oishi Y, Watanabe Y, Yoshida Y, Sato Y, Hiraishi T, et al. (2012) Hypermethylation of Sox17 gene is useful as a molecular diagnostic application in early gastric cancer. Tumour Biol 33: 383-393. [Crossref]
- 15. Baba S, Oishi Y, Watanabe Y, Oikawa R, Morita R, et al. (2011) Gastric wash-based molecular testing for antibiotic resistance in Helicobacter pylori. Digestion 84: 299-305. [Crossref]
- 16. Kahlert C, Kalluri R (2013) Exosomes in tumor microenvironment influence cancer progression and metastasis. J Mol Med (Berl) 91: 431-437. [Crossref]
- 17. Saleem SN, Abdel-Mageed AB (2014) Tumor-derived exosomes in oncogenic reprogramming and cancer progression. Cell Mol Life Sci.[Crossref]
- 18. Kosaka N, Yoshioka Y, Tominaga N, Hagiwara K, Katsuda T, et al. (2014) Dark side of the exosome: the role of the exosome in cancer metastasis and targeting the exosome as a strategy for cancer therapy. Future Oncol 10: 671-681. [Crossref]
- 19. Tanaka Y, Kamohara H, Kinoshita K, Kurashige J, Ishimoto T, et al. (2013) Clinical impact of serum exosomal microRNA-21 as a clinical biomarker in human esophageal squamous cell carcinoma. Cancer 119: 1159-1167. [Crossref]
- 20. Valadi H, Ekström K, Bossios A, Sjöstrand M, Lee JJ, et al. (2007) Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat Cell Biol 9: 654-659.[Crossref]
- 21. Taylor DD, Gercel-Taylor C (2008) MicroRNA signatures of tumor-derived exosomes as diagnostic biomarkers of ovarian cancer. Gynecol Oncol 110: 13-21. [Crossref]
- 22. Kahlert C, Melo SA, Protopopov A, Tang J, Seth S, et al. (2014) Identification of double-stranded genomic DNA spanning all chromosomes with mutated KRAS and p53 DNA in the serum exosomes of patients with pancreatic cancer. J Biol Chem 289: 3869-3875. [Crossref]
- 23. Kosaka N, Yoshioka Y, Hagiwara K, Tominaga N, Katsuda T, et al. (2013) Trash or Treasure: extracellular microRNAs and cell-to-cell communication. Front Genet 4: 173. [Crossref]
- 24. Balaj L, Lessard R, Dai L, Cho YJ, Pomeroy SL, et al. (2011) Tumour microvesicles contain retrotransposon elements and amplified oncogene sequences. Nat Commun 2: 180. [Crossref]
- 25. Guescini M, Genedani S, Stocchi V, Agnati LF (2010) Astrocytes and Glioblastoma cells release exosomes carrying mtDNA. J Neural Transm 117: 1-4. [Crossref]
- 26. Guescini M, Guidolin D, Vallorani L, Casadei L, Gioacchini AM, et al. (2010) C2C12 myoblasts release micro-vesicles containing mtDNA and proteins involved in signal transduction. Exp Cell Res 316: 1977-1984. [Crossref]
- 27. Thakur BK, Zhang H, Becker A, Matei I, Huang Y, et al. (2014) Double-stranded DNA in exosomes: a novel biomarker in cancer detection. Cell Res 24: 766-769. [Crossref]
- 28. Igarashi S, Suzuki H, Niinuma T, Shimizu H, Nojima M, et al. (2010) A novel correlation between LINE-1 hypomethylation and the malignancy of gastrointestinal stromal tumors. Clin Cancer Res 16: 5114-5123. [Crossref]
- 29. Yoshioka Y, Konishi Y, Kosaka N, Katsuda T, Kato T, et al. (2013) Comparative marker analysis of extracellular vesicles in different human cancer types. J Extracell Vesicles 2. [Crossref]
- 30. Camacho L, Guerrero P, Marchetti D (2013) MicroRNA and protein profiling of brain metastasis competent cell-derived exosomes. PLoS One 8: e73790. [Crossref]
- 31. Sinha A, Ignatchenko V, Ignatchenko A, Mejia-Guerrero S, Kislinger T (2014) In-depth proteomic analyses of ovarian cancer cell line exosomes reveals differential enrichment of functional categories compared to the NCI 60 proteome. Biochem Biophys Res Commun 445: 694–701. [Crossref]
- 32. Palazzolo G, Albanese NN, DI Cara G, Gygax D, Vittorelli ML, et al. (2012) Proteomic analysis of exosome-like vesicles derived from breast cancer cells. Anticancer Res 32: 847-860. [Crossref]
- 33. Watanabe Y, Maeda I, Oikawa R, Wu W, Tsuchiya K, et al. (2013) Aberrant DNA methylation status of DNA repair genes in breast cancer treated with neoadjuvant chemotherapy. Genes Cells 18: 1120-1130. [Crossref]
- 34. Mittra I, Nair NK, Mishra PK (2012) Nucleic acids in circulation: are they harmful to the host? J Biosci 37: 301-312. [Crossref]
- 35. Schwarzenbach H, Hoon DS, Pantel K (2011) Cell-free nucleic acids as biomarkers in cancer patients. Nat Rev Cancer11: 426-437. [Crossref]
Editorial Information
Editor-in-Chief
Masayoshi Yamaguchi
Emory University School of Medicine
Article Type
Research Article
Publication history
Received: September 15, 2014
Accepted: September 24, 2014
Published: September 29, 2014
Acknowledgements
This work was supported in part by a grants-in-aid of The Public Trust For Clinical Cancer Research (to H.Y.), Suzuken Memorial Foundation (to F.I.), Daiwa Securities Health Foundation (to F.I.), The Japanese Foundation for Research and Promotion of Endoscopy (to H.Y.), and Kobayashi Foundation for Cancer Research (to H.Y.).
Copyright
©2014 Yoshida Y. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation
Yoshida Y, Yamamoto H, Morita R, Oikawa R, Matsuo Y, et al. (2014). Detection of DNA methylation of gastric juice-derived exosomes in gastric cancer. Integr Mol Med, 1: DOI: 10.15761/IMM.1000105




