Plasma plasmalogens (Pls) may serve as potential biomarkers not only for rare peroxisomal diseases but also for general disorders related to oxidative stress and aging. Recent clinical observational studies demonstrated that low levels of plasma Pls are risk factors for atherosclerosis and dementia. Serum levels of Pls showed a strong positive correlation with high-density lipoprotein (HDL) cholesterol concentration, suggesting that Pls may be involved in metabolism or the function of HDL. Increasing the levels of plasma Pls may serve as a novel therapeutic strategy for preventing diseases associated with oxidative stress and aging. Therefore, we and other groups elevated plasma Pl levels in laboratory animals or humans through administration of myo-inositol, monounsaturated long-chain fatty acids, and the hypolipidemic agent, statin. However, their effects on the gene expression of Pl biosynthetic enzymes remain unknown. To gain insight into the manipulation of Pl biosynthesis and the relationship between Pl biosynthesis and HDL metabolism, we examined target gene expression by real time reverse transcription polymerase chain reaction (RT-PCR) in hepatoma HepG2 cells treated with various test substances. Monounsaturated long-chain fatty acids such as oleic acid and erucic acid, myo-inositol, and the Pl precursor alkylglycerol, all of which supply materials or coenzymes for Pl biosynthesis, unexpectedly reduced the expression of the genes for Pl biosynthetic enzymes. These results suggest the presence of strict regulation of Pl homeostasis. In contrast, pitavastatin induced peroxisome biogenesis and promoted the expression of peroxisomal Pl biosynthetic enzymes and HDL metabolism-associated proteins such as apoprotein A1 and ATP-binding cassette transporter A1. This was likely through enhancement of peroxisome proliferator-activated receptor (PPAR) expression. These findings suggest that there may be a physiological relationship between Pl biosynthesis and HDL metabolism via peroxisomal status.
high-density lipoprotein, human hepatocellular liver carcinoma cell line, peroxisome, plasmalogens, reverse transcription polymerase chain reaction
Abbreviations
DHA: Docosahexaenoic Acid; Far 1: Fatty Acyl CoA Reductase 1; HG: 1-O-hexadecyl-sn-glycerol; MI: myo-inositol; PlsCho: Choline Plasmalogen; PlsEtn: Ethanolamine Plasmalogen; PPARs: Peroxisome Proliferator-Activated Receptors.
Age-related diseases, such as atherosclerosis and dementia, are associated with oxidative stress and chronic inflammation [1]. Peroxisomal as well as mitochondrial dysfunction may be related to aging and age-related pathologies, possibly through the derangement of redox homeostasis [2,3]. Plasmalogens (PIs), a subclass of glycerophospholipids possessing a vinyl-ether bond at the sn-1 position, are biosynthesized and regulated in peroxisomes [4-6]. Therefore, plasma Pls may reflect the systemic functional state of peroxisomes, and serve as potential biomarkers for diseases related to oxidative stress and aging [7-9]. Human plasma Pls are synthesized mainly in the liver and secreted into the blood as lipoprotein components. To investigate the clinical significance of plasma Pls, we developed three promising analytical methods [10-13]. Our research lab and other investigators have demonstrated in clinical observational studies that low levels of plasma Pl are a risk factor for atherosclerosis and dementia [14-18]. Serum levels of Pl showed a strong positive correlation with high-density lipoprotein (HDL) cholesterol concentration [14,15], suggesting that Pls may be involved in metabolism or HDL functions.
Accordingly, we attempted to increase the levels of plasma Pls as a preventative strategy for diseases associated with oxidative stress and aging. This was achieved in laboratory animals as well as humans through administration of the Pl precursor alkylglycerol [19], myo-inositol (MI) [20,21], monounsaturated long-chain fatty acids [22], and the hypolipidemic agent, statin [23]. However, their effects on the gene expression of Pl biosynthetic enzymes remain unknown. To gain insight into the mechanisms mediating the enhancement of Pl biosynthesis, and its relationship with HDL metabolism, we examined target gene expression in HepG2 cells treated with various test substances.
Materials
Pitavastatin and simvastatin were kindly provided by Nissan Chemical Industries, Ltd. (Tokyo, Japan) and Merck Research Laboratories (Rahway, NJ), respectively. Fenofibrate was a kind gift from ASKA Pharmaceutical Co., Ltd. (Tokyo, Japan). Wy14643 was purchased from Wako Pure Chemical Industries, Ltd. (Osaka, Japan). Fatty acids such as oleic acid, erucic acid, nervonic acid, linoleic acid, arachidonic acid, eicosapentaenoic acid (EPA), docosahexaenoic acid (DHA), and β-estradiol (E2) were purchased from Sigma-Aldrich (St. Louis, MO). N-palmitoyl-D-erythro-sphingosylphosphorylcholine (SM18), N-lignoceroyl-D-erythro-sphingosylphosphorylcholine (SM24), 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine (POPC), and 1-O-hexadecyl-sn-glycerol (HG) were obtained from Avanti Polar Lipids (Alabaster, AL). Myo-inositol (MI) was a kind gift from TSUNO Co., Ltd. (Wakayama, Japan).
Cell culture and treatment with test substances
HepG2 cells (RIKEN BioResource Center, Tukuba, Ibaragi, Japan) were cultured in Dulbecco’s Modified Eagle Medium (Gibco) containing 10% fetal bovine serum (Gibco), 100 μg/mL streptomycin sodium, and 100 U/mL penicillin G sodium (Meiji Seika Pharma Co., Ltd. Tokyo, Japan) at 37°C and 5% CO2. Test substances were dissolved in distilled water, ethanol, or dimethyl sulfoxide and then passed through a membrane filter (0.45 µm) for sterilization. The test substance solution was added to the cell culture medium at a desired concentration, adjusted to a vehicle concentration of less than 0.1%. Cells were incubated with test substances at 37°C for 24 h.
Real-time RT-PCR
Total RNA was prepared using Trizol (Invitrogen, Carlsbad, CA, USA), and cDNA was synthesized from 1.0 μg RNA with GeneAmp™ RNA PCR (Applied Biosystems, Branchburg, NJ, USA) using random hexamers. Real-time RT-PCR was performed using LightCycler-FastStart DNA Master SYBR-Green 1 (Roche, Tokyo, Japan), according to the manufacturer's instructions. The reaction mixture (20 μL) contained LightCycler-FastStart DNA Master SYBR-Green 1, 4 mM MgCl2, 0.5 μM of the upstream and downstream PCR primers, and 2 mL of the first-strand cDNA as a template. The target genes and their primers are shown in Table 1. To control variations in the reactions, all PCR reactions were normalized against GAPDH or β-actin expression. The results of pitavastatin are shown as the mean ± SEM (Figure 1). Statistical analyses were performed using Stat Flex ver.6 (Artech Co. Ltd., Osaka, Japan).
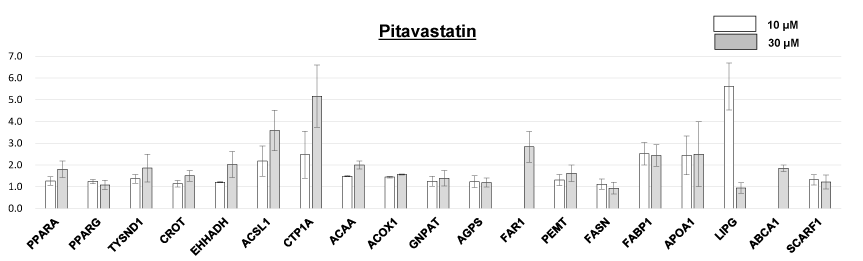
Figure 1. Effects of pitavastatin on target gene expression in HepG2 cells. Values were calculated as fold changes relative to each target mRNA expression of control cells without pitavastatin treatment. The effects of 10 uM of pitavastatin was not tested for FAR1 an ABCA1. Columns and bars represent means ± standard error (SE).
Table 1. Primers used for analysis for expression of target genes.
Gene |
Protein |
|
5’Primer 3’ |
Gene Bank accession no. |
PPARA |
peroxisome proliferator-activated receptor alpha |
Forward primer |
ATGGTGGACACGGAAAGCC |
NM-005036 |
|
|
Reverse primer |
CGATGGATTGCGAAATCTCTTGG |
|
PPARG |
peroxisome proliferator-activated receptor gamma |
Forward primer |
GGGATCAGCTCCGTGGATCT |
NM-138711 |
|
|
Reverse primer |
TGCACTTTGGTACTCTTGAAGTT |
|
TYSND1 |
trypsin domain containing 1 |
Forward primer |
TGCAGCGGGGTAATCCTGA |
NM-173555 |
|
|
Reverse primer |
CCTCCGACACTTCGTCATCC |
|
CROT |
carnitine O-octanoyltransferase |
Forward primer |
GTGGTGGCTGAATGTTGCCTA |
NM-021151 |
|
|
Reverse primer |
TTGGAGGCCAGTAGTGTTCAA |
|
EHHADH |
enoyl-CoA, hydratase/3-hydroxyacyl CoA dehydrogenase |
Forward primer |
AAACTCAGACCCGGTTGAAGA |
NM-001166415 |
|
|
Reverse primer |
TTGCAGAGTCTACGGGATTCT |
|
ACSL1 |
acyl-CoA synthetase long-chain family member 1 |
Forward primer |
GCCGAGTGGATGATAGCTGC |
NM-004457 |
|
|
Reverse primer |
ATGGCTGGACCTCCTAGAGTG |
|
CTP1A |
carnitine palmitoyltransferase 1A (liver) |
Forward primer |
TCCAGTTGGCTTATCGTGGTG |
NM-001876 |
|
|
Reverse primer |
TCCAGAGTCCGATTGATTTTTGC |
|
ACAA1 |
acetyl-Coenzyme A acyltransferase 1 |
Forward primer |
GCGGTTCTCAAGGACGTGAAT |
NM-001607 |
|
|
Reverse primer |
GTCTCCGGGATGTCACTCAGA |
|
ACOX1 |
acyl-Coenzyme A oxidase 1, palmitoyl |
Forward primer |
ACTCGCAGCCAGCGTTATG |
NM-007292 |
|
|
Reverse primer |
AGGGTCAGCGATGCCAAAC |
|
GNPAT |
glyceronephosphate O-acyltransferase |
Forward primer |
GAGGAGGCATGTCAGTGACTT |
NM-014236 |
|
|
Reverse primer |
ACAAAACCGAATGGCTCCAAG |
|
AGPS |
alkylglycerone phosphate synthase |
Forward primer |
TGAGTACCAATGAGTGCAAAGC |
NM-003659 |
|
|
Reverse primer |
GGTAAACCCATGCCACTAAGAG |
|
FAR1 |
fatty acyl-CoA reductase 1: |
Forward primer |
AGACACCACAAGAGCGAGTG |
NM-032228 |
|
|
Reverse primer |
CCAGTTTAGGTTGGGTGAGTTC |
|
PEMT |
phosphatidylethanolamine N-methyltransferase |
Forward primer |
CTGGAATGTGGTTGCACGATG |
NM-148172 |
|
|
Reverse primer |
GCTTAGAGAGTAGCAGGCCA |
|
FASN |
fatty acid synthase |
Forward primer |
AAGGACCTGTCTAGGTTTGATGC |
NM-004104 |
|
|
Reverse primer |
TGGCTTCATAGGTGACTTCCA |
|
FABP1 |
fatty acid binding protein 1 (liver) |
Forward primer |
ATGAGTTTCTCCGGCAAGTACC |
NM-001443 |
|
|
Reverse primer |
CTCTTCCGGCAGACCGATTG |
|
APOA1 |
apolipoprotein A1 |
Forward primer |
CCCTGGGATCGAGTGAAGGA |
NM-000039 |
|
|
Reverse primer |
CTGGGACACATAGTCTCTGCC |
|
LIPG |
lipase, endothelial |
Forward primer |
GGGAGCCCCGTACCTTTTG |
NM-006033 |
|
|
Reverse primer |
CCTCACAGATGGTTTGACCTCA |
|
ABCA1 |
ATP-binding cassette, sub-family A1 |
Forward primer |
GGAAGAACAGTCATTGGGACAC |
NM-080282 |
|
|
Reverse primer |
GCTACAAACCCTTTTAGCCAGT |
|
SCARF1 |
scavenger receptor class B, member 1 (SR-B1) |
Forward primer |
CCGATCAGACCTCAAGGACAG |
NM-145352 |
|
|
Reverse primer |
CCCAGGGTAGCTTGTGGGA |
|
GAPDH |
glyceraldehyde-3-phosphate dehydrogenase |
Forward primer |
GGAGCGAGATCCCTCCAAAAT |
NM-001256799 |
|
|
Reverse primer |
GGCTGTTGTCATACTTCTCATGG |
|
β-Actin |
beta actin |
Forward primer |
CATGTACGTTGCTATCCAGGC |
NM-001101 |
|
|
Reverse primer |
CTCCTTAATGTCACGCACGAT |
|
Effect of test substances on target gene expression in HepG2 cells
The effects of several test substances on target mRNA expression in HepG2 cells were examined by real time RT-PCR (Table 2). The target genes were chosen such as to cover more or less the enzymes involved in Pl synthesis, functions of peroxisome and lipid metabolism. Pitavastatin and simvastatin enhanced the expression of genes for apolipoprotein A1 (gene name: APOA1), ATP-binding cassette A1 (ABCA1), and fatty acid binding protein 1 (liver) (FABP1), as well as peroxisomal β-oxidation enzymes, such as acyl-CoA synthetase long-chain family member 1 (ACSL1) and carnitine palmitoyltransferase 1A (liver) (CTP1A). Peroxisome proliferator-activated receptor (PPAR) agonists, fenofibrate and Wy14643, also enhanced ACSL1 and CTP1A expression. These agonists reduced the expression of genes for other peroxisomal β-oxidation enzymes, such as enoyl-CoA, hydratase/3-hydroxyacyl CoA dehydrogenase (EHHADH) and acetyl-Coenzyme A acyltransferase 1 (ACAA1), as well as FABP1 expression. 1-Palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine (POPC), a proposed endogenous ligand for PPAR alpha [24], increased APOA1, phosphatidylethanolamine N-methyltransferase (PEMT), fatty acid synthase (FASN), and lipase, endothelial (LIPG) expression. Monounsaturated long-chain fatty acids such as oleic acid (C18:1) and erucic acid (C22:1) increased APOA1, PEMT, and FASN expression and decreased fatty acyl-CoA reductase 1 (FAR1) and FABP1 expression, while nervonic acid (C24:1) exhibited no significant effects on target gene expression. However, polyunsaturated fatty acids had different effects on gene expression. For instance, linoleic acid (C18:2) and arachidonic acid (C20:4) increased ACSL1 expression and lowered FABP1 expression. EPA (C22:5) also increased APOA1 expression and DHA (C22:6) increased FAR1 and ABCA1 expression. MI treatment resulted in decreased expression of multiple genes such as APOA1, PEMT, and FASN, as well as Pl biosynthetic enzymes such as glyceronephosphate O-acyltransferase (GNPAT) and alkylglycerone phosphate synthase (AGPS). MI also reduced the expression of peroxisomal β-oxidation enzymes such as carnitine O-octanoyltransferase (CROT), ACSL1, and ACAA1. 1-O-hexadecyl-sn-glycerol (HG), a precursor for Pl biosynthesis [25], lowered the expression of GNPAT and FAR1, as well as FASN and LIPG. Sphingomyelins (SM18 and SM24) and β-estradiol (E2) had no significant effects on target gene expression.
Table 2. Effects of test substances on target gene expression in HepG2 cells.
Wy14643: Agonist of PPARα and PPARγ; POPC: 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine; EPA: Eicosapentaenoic acid; DHA: Docosahexaenoic acid; SM18: Sphingomyelin containing stearic acid; SM24: Sphingomyelin containing lignoceric acid; E2: β-estradiol; MI: Myo-inositol; HG: 1-o-hexadecyl-sn-glycerol.
Test substances |
Pitavastatin |
Simvastatin |
Fenofibrate |
Wy14643 |
POPC |
Oleic acid |
Erucic acid |
Nervonic acid |
Linoleic acid |
Arachidonic acid |
EPA |
DHA |
SM18 |
SM24 |
E2 |
MI |
HG |
|
|
|
|
|
|
|
|
|
C18:1 |
C22:1 |
C24:1 |
C18:2 |
C20:4 |
C20:5 |
C22:6 |
|
|
|
|
|
|
|
|
Concentrations |
10-100 µM |
10-30 µM |
10-100 µM |
10-100 µM |
50-250 µM |
10-50 µM |
10-50 µM |
10-100 µM |
10-100 µM |
10-100 µM |
10-100 µM |
10-100 µM |
10-50 µM |
10-50 µM |
1-100 nM |
0.2-5 mM |
60 µM |
|
|
|
Number of tests |
5 times |
Twice |
Twice |
Once |
Once |
5 times |
5 times |
Twice |
Twice |
Twice |
Twice |
3 times |
5 times |
5 times |
Twice |
Once |
Once |
|
|
|
Gene name |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
++ |
Obvious rise (more than 2-fold) |
PPARA |
+ |
|
± |
± |
± |
± |
± |
± |
± |
± |
± |
± |
|
|
± |
± |
± |
|
+ |
Moderate rise (more than 1.5-fold) |
PPARG |
± |
|
± |
± |
|
± |
± |
± |
± |
± |
± |
± |
|
|
± |
± |
± |
|
± |
No change |
TYSND1 |
+ |
|
± |
± |
|
± |
± |
± |
± |
± |
± |
± |
± |
± |
± |
± |
± |
|
- |
Moderate descent (less than 0.75) |
CROT |
+ |
|
± |
- |
|
+ |
± |
± |
± |
± |
± |
+ |
± |
± |
± |
- |
± |
|
-- |
Obvious descent (less than 0.5) |
EHHADH |
+ |
|
-- |
- |
|
± |
± |
± |
± |
- |
± |
± |
|
|
± |
± |
± |
|
|
|
ACSL1 |
++ |
+ |
++ |
++ |
± |
± |
± |
± |
++ |
+ |
± |
± |
|
|
± |
- |
|
|
|
|
CTP1A |
++ |
+ |
++ |
++ |
-- |
± |
± |
± |
± |
± |
± |
± |
± |
± |
± |
++ |
++ |
|
|
|
ACAA1 |
+ |
|
- |
- |
|
± |
± |
± |
± |
± |
± |
± |
± |
± |
± |
- |
|
|
|
|
ACOX1 |
+ |
|
± |
± |
|
± |
± |
± |
± |
± |
± |
± |
|
|
± |
± |
|
|
|
|
GNPAT |
+ |
± |
± |
- |
± |
± |
± |
± |
± |
± |
± |
± |
± |
± |
± |
- |
- |
|
|
|
AGPS |
+ |
± |
± |
± |
± |
± |
± |
± |
± |
± |
± |
± |
± |
± |
± |
- |
± |
|
|
|
FAR1 |
++ |
± |
± |
|
|
- |
- |
± |
± |
± |
± |
+ |
|
|
|
|
- |
|
|
|
PEMT |
+ |
± |
± |
± |
+ |
+ |
+ |
± |
± |
± |
± |
± |
± |
± |
± |
-- |
+ |
|
|
|
FASN |
± |
± |
± |
± |
++ |
+ |
+ |
|
|
|
|
|
± |
± |
± |
-- |
-- |
|
|
|
FABP1 |
++ |
++ |
-- |
- |
|
- |
- |
± |
- |
- |
± |
± |
|
|
± |
+ |
± |
|
|
|
APOA1 |
++ |
+ |
± |
± |
+ |
++ |
++ |
± |
± |
± |
+ |
± |
± |
± |
± |
-- |
± |
|
|
|
LIPG |
++ |
|
|
|
++ |
± |
± |
|
|
|
|
|
± |
± |
± |
± |
- |
|
|
|
ABCA1 |
+ |
+ |
± |
|
|
± |
± |
± |
± |
± |
± |
+ |
|
|
|
± |
|
|
|
|
SCARF1 |
± |
|
|
|
± |
± |
± |
|
|
|
|
|
± |
± |
± |
± |
± |
|
|
|
Effects of pitavastatin on target gene expression in HepG2 cells
Pitavastatin enhanced the expression of multiple genes (Figure 1). Increased expression of PPAR alpha (PPARA) and trypsin domain containing 1 (TYSND1) [26], as well as peroxisomal β-oxidation enzymes were observed with pitavastatin treatment. Pitavastatin also enhanced the expression of HDL metabolism-associated proteins such as APOA1, ABCA1, and LIPG by greater than two-fold. Furthermore, pitavastatin enhanced the expression of Pl biosynthetic enzymes such as GNPAT, AGPS, and FAR1. Particularly, the expression of the rate-limiting enzyme of Pl biosynthesis, FAR1, was increased by nearly 3-fold. In addition, pitavastatin augmented by approximately 1.5-fold the expression of PEMT, which specifically localizes in the liver [27] and possibly participates in the conversion of ethanolamine plasmalogen (PlsEtn; 1-O-alk-1’-enyl-2-acyl-sn-glycero-3-phosphoethanolamine) to choline plasmalogen (PlsCho; 1-O-alk-1’-enyl-2-acyl-sn-glycero-3-phosphocholine) [28].
Monounsaturated long-chain fatty acids such as oleic acid (C18:1) and erucic acid (C22:1), myo-inositol, and Pl precursors, alkylglycerol and HG, have been reported to increase Pl levels in laboratory animals and humans [19-23]. However, they unexpectedly reduced the gene expression of Pl biosynthetic enzymes in HepG2 cells (Table 2). Monounsaturated long-chain fatty acids are preferred substrates for peroxisomal β-oxidation, and the resulting acetyl CoA is preferentially utilized for the synthesis of ether phospholipids including Pls [29,30]. The decreased expression of FAR1 in HepG2 cells treated with C18:1 and C22:1 may have resulted from the negative feedback from the overproduction of Pls. MI is presumed to enhance Pl biosynthesis through NADPH generation during MI catabolism [31], since Far 1 is activated via NADPH binding [32]. Therefore, the suppressed expression of Pl biosynthetic enzymes in cells treated with MI could also be caused by the negative feedback from overproduction of Pl. Similarly, the reduced expression of Pl biosynthetic enzymes in cells treated with HG was thought to be attributed to overload of Pls in the cells.
However, DHA and pitavastatin increased the gene expression of Pl biosynthetic enzymes in HepG2 cells (Table 2, Figure 1). Because DHA is preferentially incorporated into Pls at the sn-2 position and Pls may function as reservoirs for these biologically active lipid mediators [33], DHA supplementation was considered to potentiate FAR1 expression, the enzyme fatty acyl CoA reductase 1 (Far 1) supplies the fatty alcohols used in the formation of ether-linked alkyl bonds. Pitavastatin, but not simvastatin, facilitated the gene expression of peroxisomal Pl biosynthetic enzymes such as GNPAT, AGPS, and FAR1. The first two steps in Pl biosynthesis, which are catalyzed by the enzymes encoded by GNPAT and AGPS, exclusively occur in peroxisomes [4,34]. In addition, the rate-limiting enzyme of Pl biosynthesis, is also peroxisomal [5]. Pitavastatin further enhanced the expression of peroxisomal PPARA and TYSND1, as well as β-oxidation enzymes (Table 2, Figure 1). This suggests that pitavastatin may increase Pl biosynthesis by facilitating peroxisome biogenesis. In addition, pitavastatin increased the expression of PEMT, which may be involved in the conversion of PlsEtn to PlsCho. Our clinical observational studies indicated that serum levels of Pls, particularly PlsCho were significantly but negatively associated with diverse risk factors for metabolic syndrome and/or atherosclerosis. Furthermore, PlsCho showed the stronger positive correlation with HDL cholesterol concentration than PlsEtn [14,15]. Pitavastatin is a strong HMG-CoA reductase inhibitor and is more potent than other statins in lowering serum total cholesterol, low-density lipoprotein cholesterol, and triglycerides with modest elevation of HDL cholesterol [35]. Recently, pitavastatin was reported to increase Pl content in HDL particles in relation to improving HDL functionality [36]. Moreover, pitavastatin promoted the expression of HDL metabolism-associated proteins such as APOA1. It is proposed that this is probably via enhancement of PPAR expression, since APOA1 and ABCA1 expression are up-regulated by PPAR agonists [37].
In conclusion, the supply of materials or coenzymes for Pl biosynthesis such as acetyl CoA derived from peroxisomal β-oxidation of monounsaturated long-chain fatty acids, the Pl precursor alkylglycerol, and NADPH from MI catabolism, suppressed the expression of Pl biosynthetic enzymes (Table 2). These results suggest that Pl homeostasis is strictly regulated, and the supplementation of these materials may be effective in restoring normal levels of Pls in Pl-deficient individuals. Since peroxisome biogenesis induced by treatment with pitavastatin promoted both the gene expression of Pl biosynthetic enzymes and HDL metabolism-associated proteins, there may be a close relationship between them as a result of their peroxisomal status. Our findings of the strong association between the serum levels of Pls, especially PlsCho and HDL-cholesterol concentration, in clinical observational studies [14,15] might reflect their regulatory gene expression levels. Furthermore, their physiological association may extend to HDL functionality, specifically, Pls may induce atheroprotective effects of HDL, such as cholesterol efflux capacity, anti-inflammatory and antioxidant activities, and endothelial protection [36,38,39].
R.M. contributed conception of the work and drafting the article.
S.N. contributed data collection.
R.M. and S.N. contributed data analysis and interpretation and final approval of the version to be published.
The authors have no conflicts of interest to report.
We are very grateful for the assistance of our colleagues from the Department of Biochemistry, Teikyo University School of Medicine.
- Chung HY, Sung B, Jung KJ, Zou Y, Yu BP (2006) The molecular inflammatory process in aging. Antioxid Redox Signal 8: 572-581. [Crossref]
- Giordano CR, Terlecky SR (2012) Peroxisomes, cell senescence, and rates of aging. Biochim Biophys Acta 1822: 1358-1362. [Crossref]
- Lavrovsky Y, Chatterjee B, Clark RA, Roy AK (2000) Role of redox-regulated transcription factors in inflammation, aging and age-related diseases. Exp Gerontol 35: 521-532. [Crossref]
- Waterham HR, Ferdinandusse S2, Wanders RJ2 (2016) Human disorders of peroxisome metabolism and biogenesis. Biochim Biophys Acta 1863: 922-933. [Crossref]
- Honsho M, Asaoku S, Fujiki Y (2010) Posttranslational regulation of fatty acyl-CoA reductase 1, Far1, controls ether glycerophospholipid synthesis. J Biol Chem 285: 8537-8542. [Crossref]
- Lessig J, Fuchs B (2009) Plasmalogens in biological systems: their role in oxidative processes in biological membranes, their contribution to pathological processes and aging and plasmalogen analysis. Curr Med Chem 16: 2021-2041. [Crossref]
- Zoeller RA, Lake AC, Nagan N, Gaposchkin DP, Legner MA, et al. (1999) Plasmalogens as endogenous antioxidants: somatic cell mutants reveal the importance of the vinyl ether. Biochem J 338 : 769-776. [Crossref]
- Nagan N, Zoeller RA (2001) Plasmalogens: biosynthesis and functions. Prog Lipid Res 40: 199-229. [Crossref]
- Braverman NE, Moser AB (2012) Functions of plasmalogen lipids in health and disease. Biochim Biophys Acta 1822: 1442-1452. [Crossref]
- Maeba R, Ueta N (2004) Determination of choline and ethanolamine plasmalogens in human plasma by HPLC using radioactive triiodide (1-) ion (125I3-). Anal Biochem 331: 169-176. [Crossref]
- Maeba R, Yamazaki Y, Nezu T, Okazaki T (2012) Improvement and validation of 125I-high-performance liquid chromatography method for determination of total human serum choline and ethanolamine plasmalogens. Ann Clin Biochem 49: 86-93. [Crossref]
- Nishimukai M, Yamashita M, Watanabe Y, Yamaz2021 Copyright OAT. All rights reservc absorption of choline plasmalogen is much higher than that of ethanolamine plasmalogen in rats. Eur J Nutr 50: 427-436. [Crossref]
- Sakasegawa SI, Maeba R, Murayama K, Matsumoto H, Sugimori D (2016) Hydrolysis of plasmalogen by phospholipase A1 from Streptomyces albidoflavus for early detection of dementia and arteriosclerosis. Biotechnol Lett 38: 109-116. [Crossref]
- Nishimukai M, Maeba R, Yamazaki Y, Nezu T, Sakurai T, et al. (2014) Serum choline plasmalogens, particularly those with oleic acid in sn-2, are associated with proatherogenic state. J Lipid Res 55: 956-965. [Crossref]
- Nishimukai M, Maeba R, Ikuta A, Asakawa N, Kamiya K, et al. (2014) Serum choline plasmalogens-those with oleic acid in sn-2-are biomarkers for coronary artery disease. Clin Chim Acta 437C: 147-154. [Crossref]
- Sutter I, Klingenberg R, Othman A, Rohrer L, Landmesser U, et al. (2016) Decreased phosphatidylcholine plasmalogens - A putative novel lipid signature in patients with stable coronary artery disease and acute myocardial infarction. Atherosclerosis 246: 130-140. [Crossref]
- Goodenowe DB, Cook LL, Liu J, Lu Y, Jayasinghe DA, et al. (2007) Peripheral ethanolamine plasmalogen deficiency: a logical causative factor in Alzheimer's disease and dementia. J Lipid Res 48: 2485-2498. [Crossref]
- Maeba R, Araki A, Ishii K, Ogawa K, Tamura Y, et al. (2016) Serum ethanolamine plasmalogens improve detection of cognitive impairment among elderly with high excretion levels of urinary myo-inositol: A cross-sectional study. Clin Chim Acta 453: 134-140. [Crossref]
- Wood PL, Smith T, Lane N, Khan MA, Ehrmantraut G, et al. (2011) Oral bioavailability of the ether lipid plasmalogen precursor, PPI-1011, in the rabbit: a new therapeutic strategy for Alzheimer's disease. Lipids Health Dis 10: 227. [Crossref]
- Maeba R, Hara H, Ishikawa H, Hayashi S, Yoshimura N, et al. (2008) Myo-inositol treatment increases serum plasmalogens and decreases small dense LDL, particularly in hyperlipidemic subjects with metabolic syndrome. J Nutr Sci Vitaminol (Tokyo) 54: 196-202. [Crossref]
- Hoffman-Kuczynski B, Reo NV (2005) Administration of myo-inositol plus ethanolamine elevates phosphatidylethanolamine plasmalogen in the rat cerebellum. Neurochem Res 30: 47-60. [Crossref]
- Yamazaki Y, Kondo K, Maeba R, Nishimukai M, Nezu T, et al. (2014) Proportion of nervonic acid in serum lipids is associated with serum plasmalogen levels and metabolic syndrome. J Oleo Sci 63: 527-537. [Crossref]
- Khan M, Singh J, Singh I (2008) Plasmalogen deficiency in cerebral adrenoleukodystrophy and its modulation by lovastatin. J Neurochem 106: 1766-1779. [Crossref]
- Wolf G (2010) Endogenous ligand for an orphan receptor. Nutr Rev 68: 316-318. [Crossref]
- Zheng H, Duclos RI Jr, Smith CC, Farber HW, Zoeller RA (2006) Synthesis and biological properties of the fluorescent ether lipid precursor 1-O-[9'-(1''-pyrenyl)] nonyl-sn-glycerol. J Lipid Res 47: 633-642. [Crossref]
- Mizuno Y, Ninomiya Y, Nakachi Y, Iseki M, Iwasa H, et al. (2013) Tysnd1 deficiency in mice interferes with the peroxisomal localization of PTS2 enzymes, causing lipid metabolic abnormalities and male infertility. PLoS Genet 9: e1003286. [Crossref]
- Vance DE, Walkey CJ, Cui Z (1997) Phosphatidylethanolamine N-methyltransferase from liver. Biochim Biophys Acta 1348: 142-150. [Crossref]
- Morita SY, Takeuchi A, Kitagawa S (2010) Functional analysis of two isoforms of phosphatidylethanolamine N-methyltransferase. Biochem J 432: 387-398. [Crossref]
- Hayashi H, Oohashi M (1995) Incorporation of acetyl-CoA generated from peroxisomal beta-oxidation into ethanolamine plasmalogen of rat liver. Biochim Biophys Acta 1254: 319-325. [Crossref]
- Hayashi H, Hara M (1997) 1-Alkenyl group of ethanolamine plasmalogen derives mainly from de novo-synthesized fatty alcohol within peroxisomes, but not extraperoxisomal fatty alcohol or fatty acid. J Biochem 121: 978-983. [Crossref]
- Pettegrew JW, Panchalingam K, Levine J, McClure RJ, Gershon S, et al. (2001) Chronic myo-inositol increases rat brain phosphatidylethanolamine plasmalogen. Biol Psychiatry 49: 444-453. [Crossref]
- Honsho M, Asaoku S, Fukumoto K, Fujiki Y (2013) Topogenesis and homeostasis of fatty acyl-CoA reductase 1. J Biol Chem 288: 34588-34598. [Crossref]
- Farooqui AA, Horrocks LA (2001) Plasmalogens, phospholipase A2, and docosahexaenoic acid turnover in brain tissue. J Mol Neurosci 16: 263-272. [Crossref]
- Ofman R, Lajmir S, Wanders RJ (2001) Etherphospholipid biosynthesis and dihydroxyactetone-phosphate acyltransferase: resolution of the genomic organization of the human gnpat gene and its use in the identification of novel mutations. Biochem Biophys Res Commun 281: 754-760. [Crossref]
- Chapman MJ (2011) Pitavastatin: novel effects on lipid parameters. Atheroscler Suppl 12: 277-284. [Crossref]
- Orsoni A, Thérond P, Tan R, Giral P, Robillard P, et al. (2016) Statin action enriches HDL3 in polyunsaturated phospholipids and plasmalogens and reduces LDL-derived phospholipid hydroperoxides in atherogenic mixed dyslipidemia. J Lipid Res 57: 2073-2087. [Crossref]
- Miller M, Rhyne J, Hamlette S, Birnbaum J, Rodriguez A (2003) Genetics of HDL regulation in humans. Curr Opin Lipidol 14: 273-279. [Crossref]
- Mandel H, Sharf R, Berant M, Wanders RJ, Vreken P, et al. (1998) Plasmalogen phospholipids are involved in HDL-mediated cholesterol efflux: insights from investigations with plasmalogen-deficient cells. Biochem Biophys Res Commun 250: 369-373. [Crossref]
- Sutter I, Velagapudi S, Othman A, Riwanto M, Manz J, et al. (2015) Plasmalogens of high-density lipoproteins (HDL) are associated with coronary artery disease and anti-apoptotic activity of HDL. Atherosclerosis 241: 539-546. [Crossref]
Editorial Information
Editor-in-Chief
Kazuhisa Nishizawa
Teikyo University
Article Type
Research Article
Publication history
Received date: January 07, 2017
Accepted date: January 31, 2017
Published date: February 03, 2017
Copyright
© 2017 Maeba R. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation
Maeba R, Nakahara S (2017) Extrinsic effectors regulating genes for plasmalogen biosynthetic enzymes in HepG2 cells. Biomed Res Clin Prac 2: DOI: 10.15761/BRCP.1000128
Corresponding author
Ryouta Maeba
Department of Biochemistry, Teikyo University School of Medicine, 2-11-1 Kaga, Itabashi-ku, Tokyo 173-8605, Japan, Tel: +81-3-3964-1211;

